Cancer Res Treat.
2018 Oct;50(4):1396-1417. 10.4143/crt.2017.537.
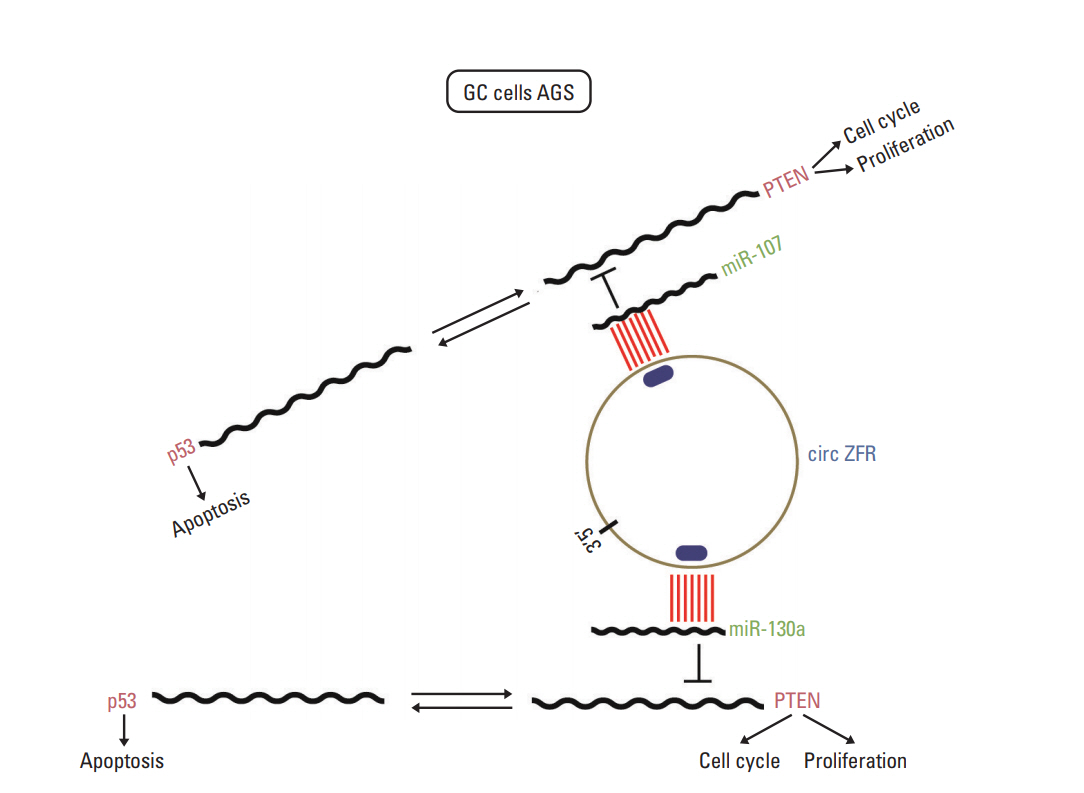
Circular RNA-ZFR Inhibited Cell Proliferation and Promoted Apoptosis in Gastric Cancer by Sponging miR-130a/miR-107 and Modulating PTEN
- Affiliations
-
- 1Department of Gastrointestinal Surgery, First Affiliated Hospital of Kunming Medical University, Kunming, China. xiufengling_njfy@163.com, ldjslyy01@163.com
- 2Department of Ultrasonics, First Affiliated Hospital of Kunming Medical University, Kunming, China.
- KMID: 2424811
- DOI: http://doi.org/10.4143/crt.2017.537
Abstract
- PURPOSE
This study aimed to probe into the associations among circular RNA ZFR (circ-ZFR), miR-130a/miR-107, and PTEN, and to investigate the regulatory mechanism of circ-ZFR-miR-130a/miR-107-PTEN axis in gastric cancer (GC).
MATERIALS AND METHODS
GSE89143 microarray data used in the study were acquired from publicly available Gene Expression Omnibus database to identify differentially expressed circular RNAs inGC tissues. The expressions of circ-ZFR, miR-130a, miR-107, and PTEN were examined by real-time reverse transcription polymerase chain reaction, while PTEN protein expression was measured by western blot. The variation of GC cell proliferation and apoptosis was confirmed by cell counting kit-8 assay and flow cytometry analysis. The targeted relationships among circZFR, miR-130a/miR-107, and PTEN were predicted via bioinformatics analysis and demonstrated by dual-luciferase reporter assay and RNA immunoprecipitation assay. The impact of ZFR on gastric tumor was further verified in xenograft mice model experiment.
RESULTS
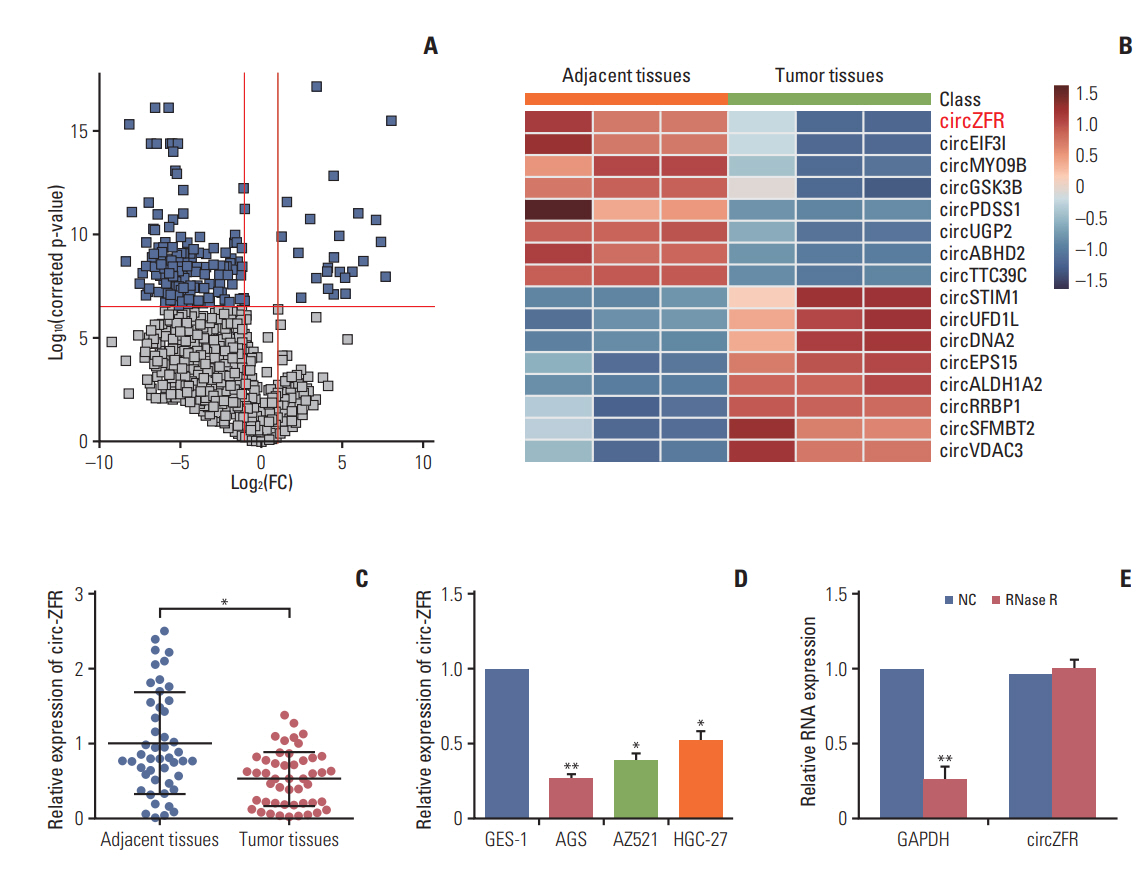
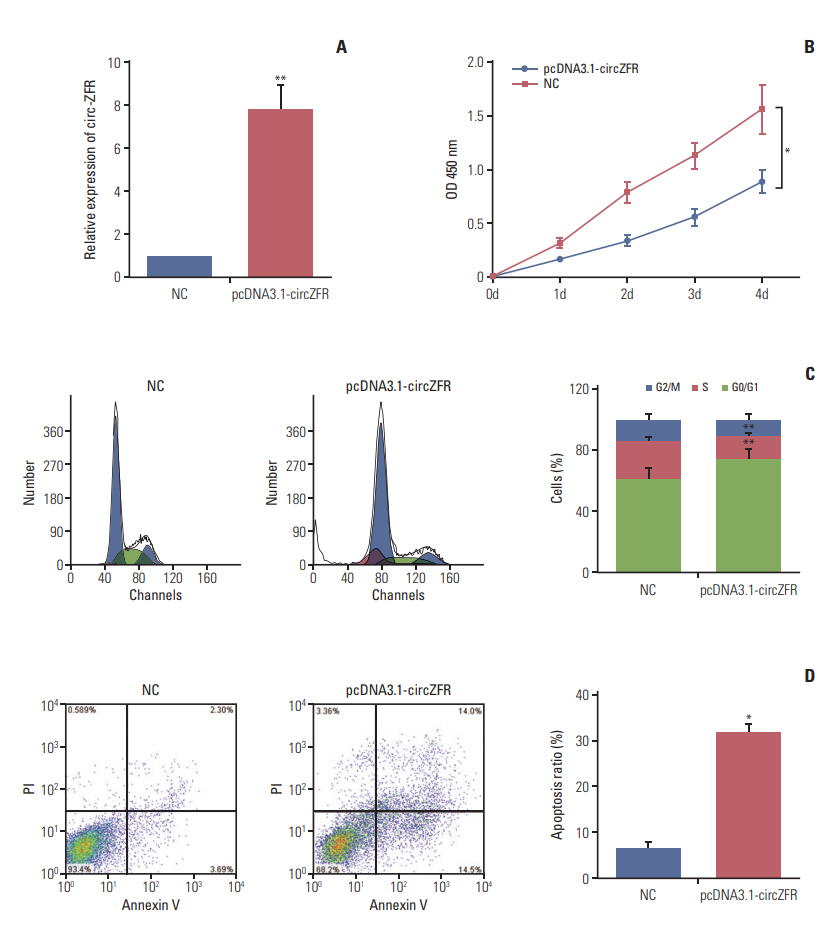
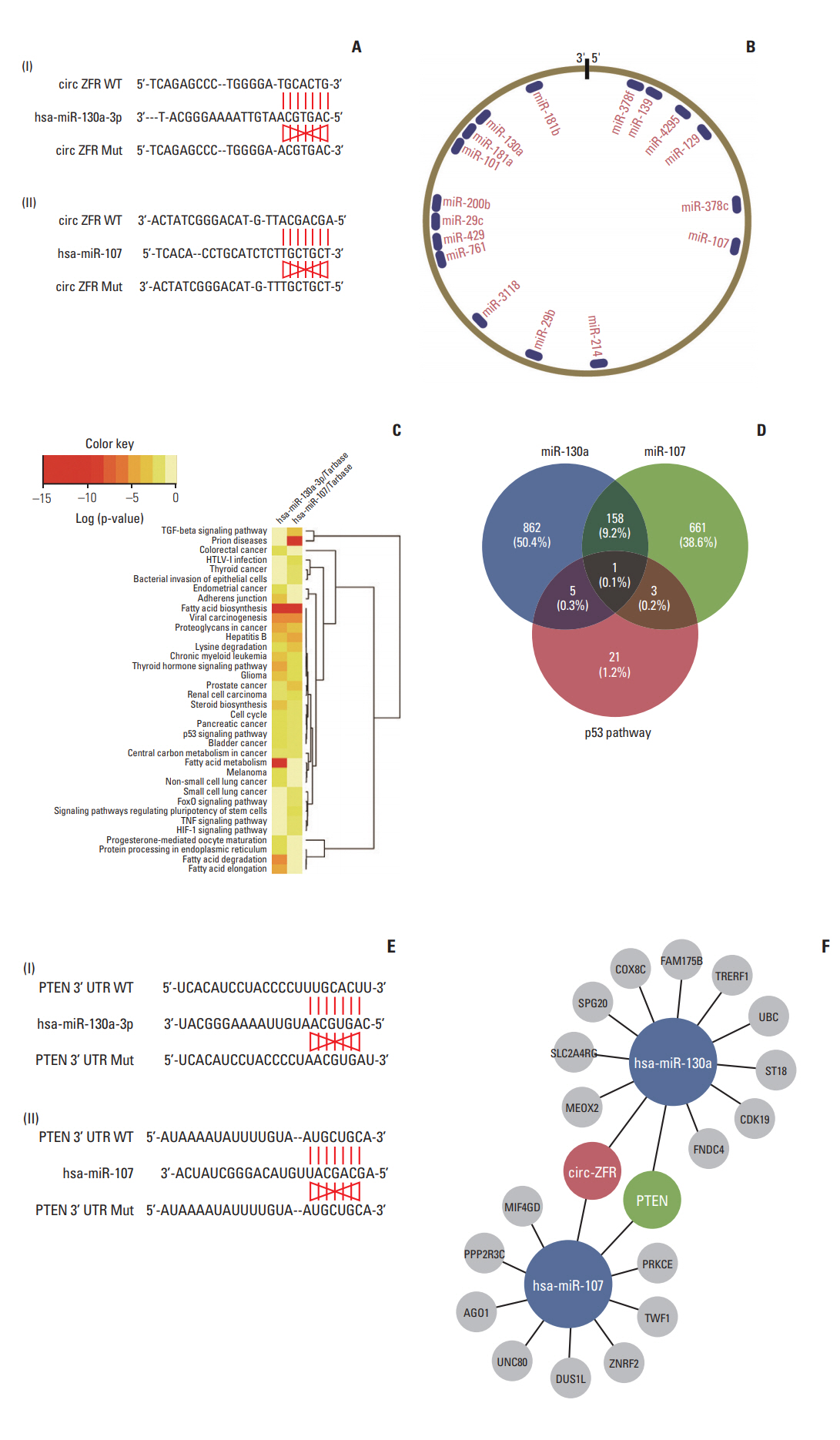
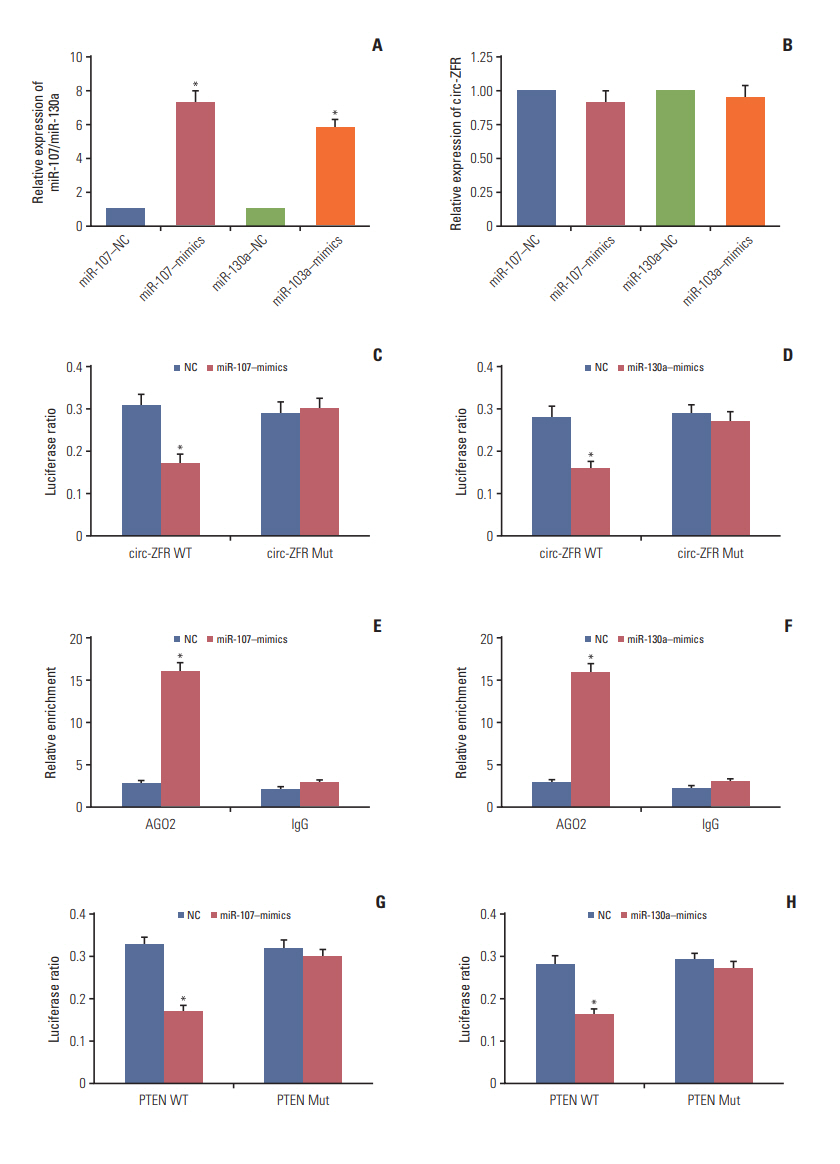
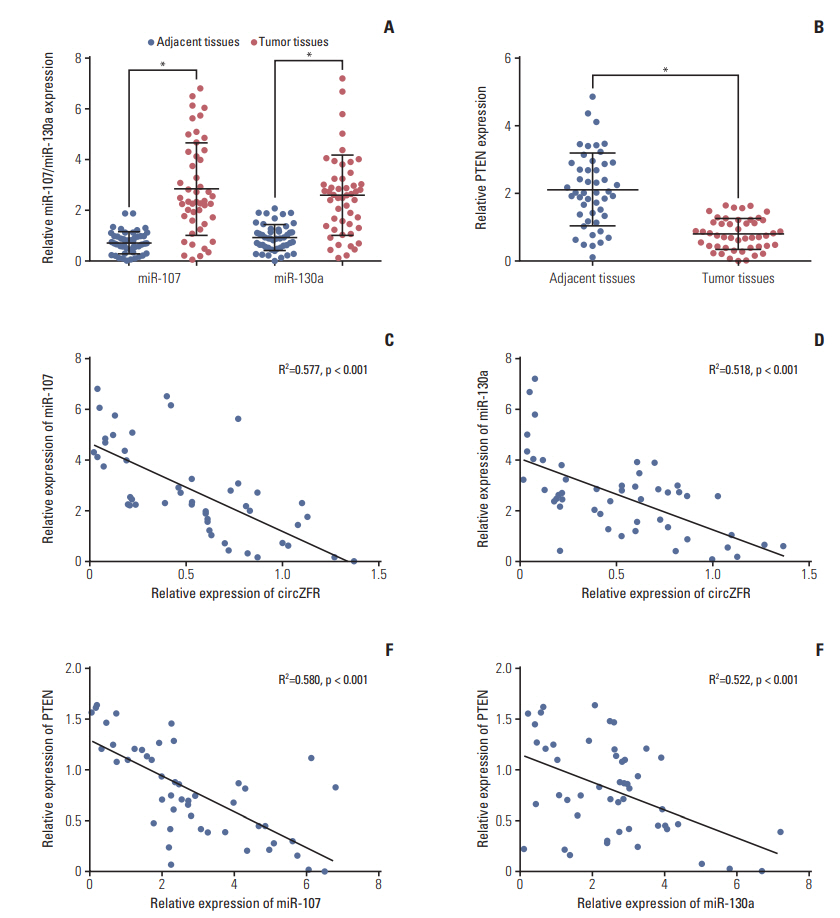
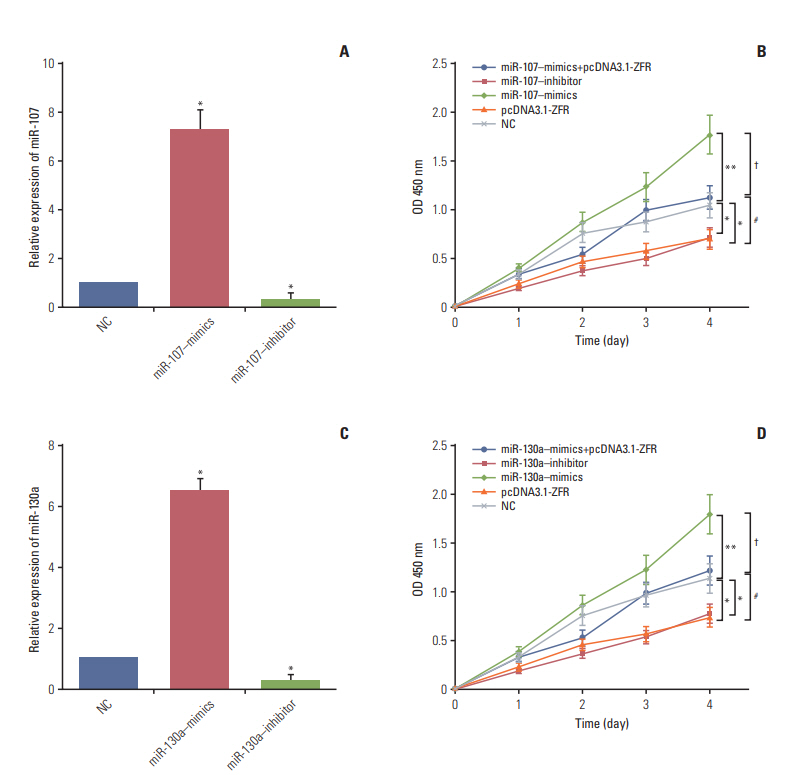
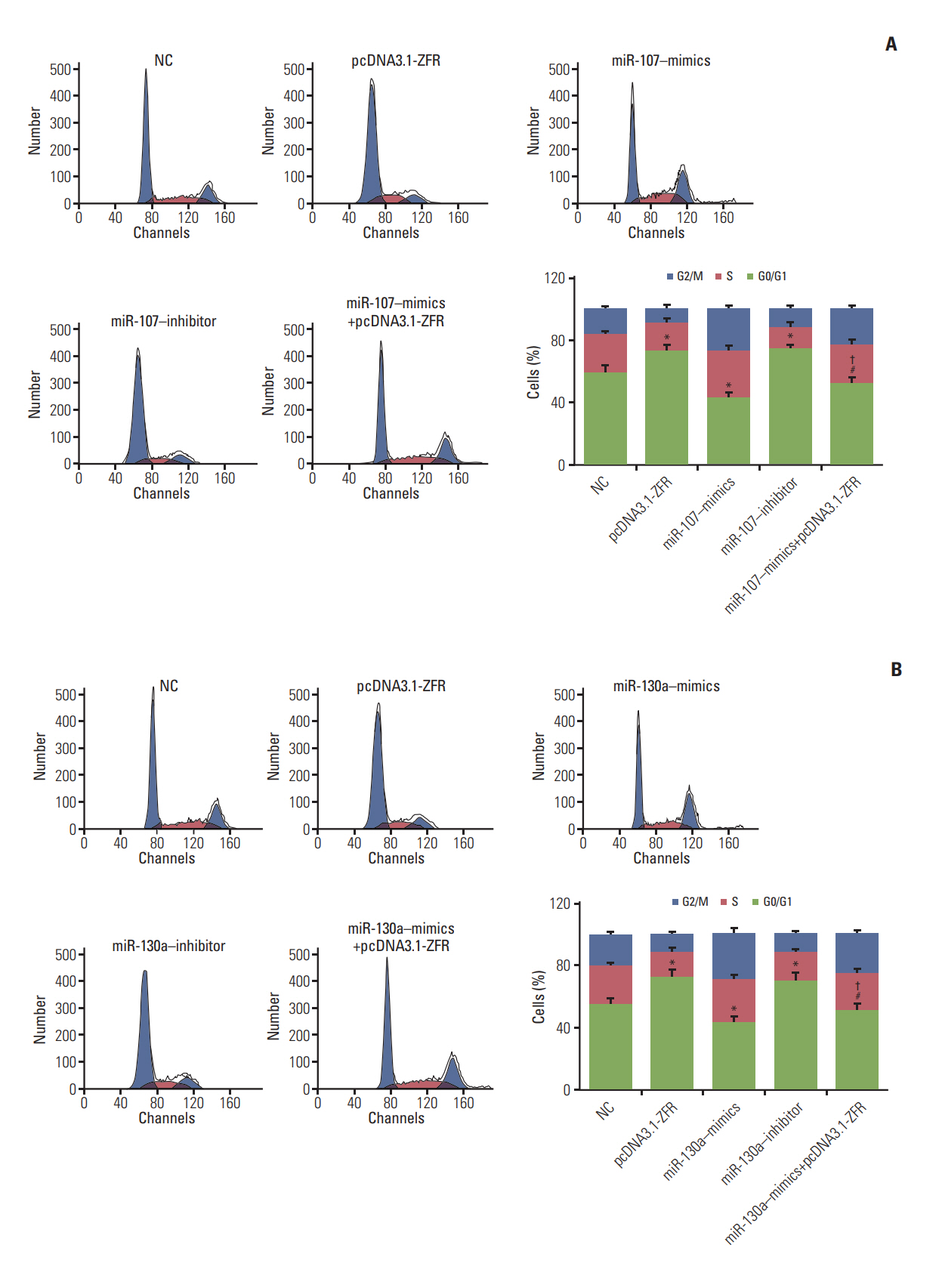
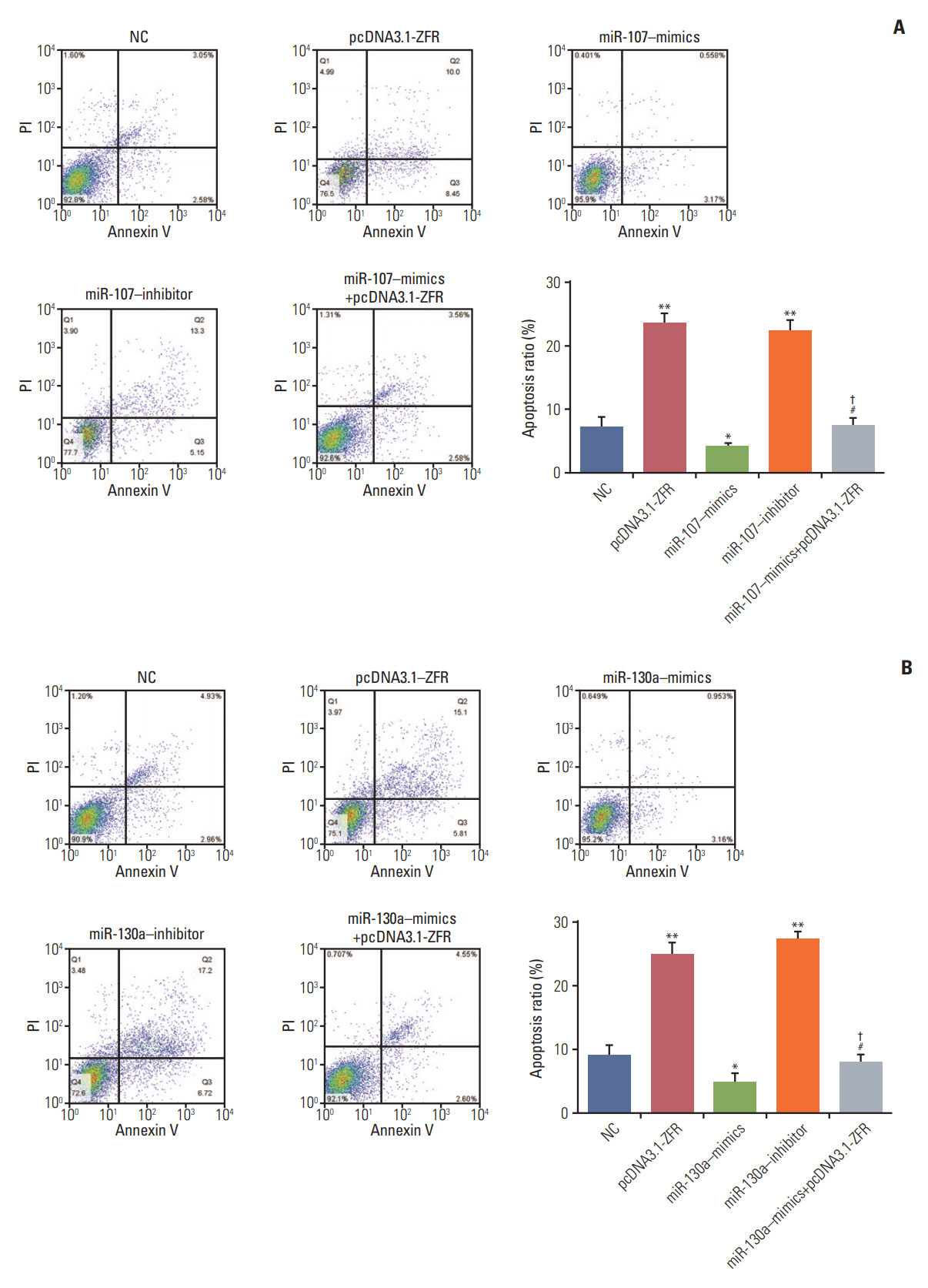
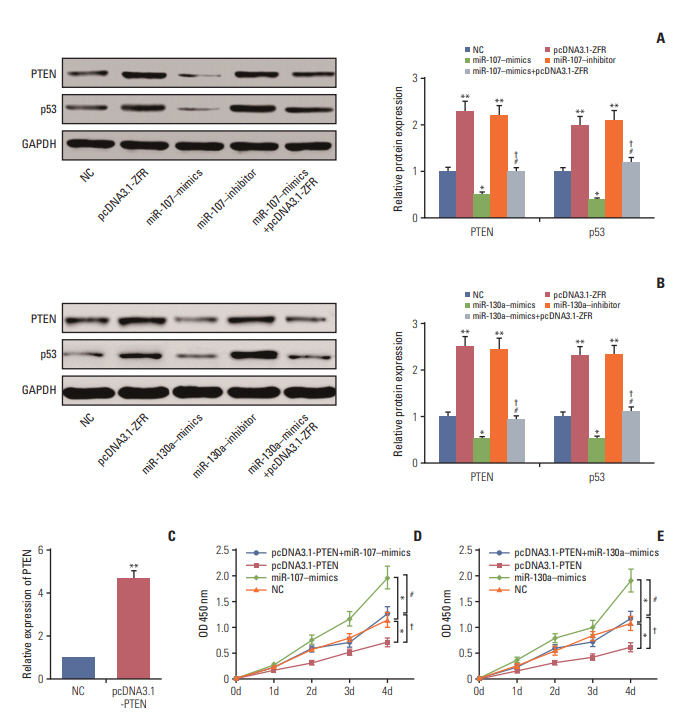
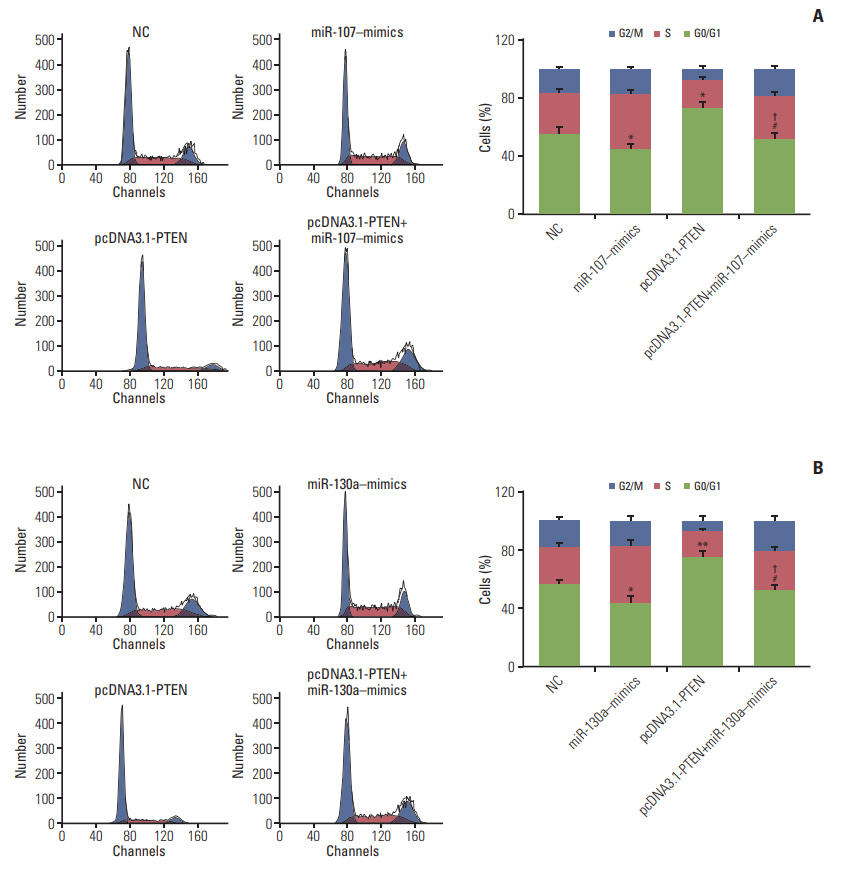
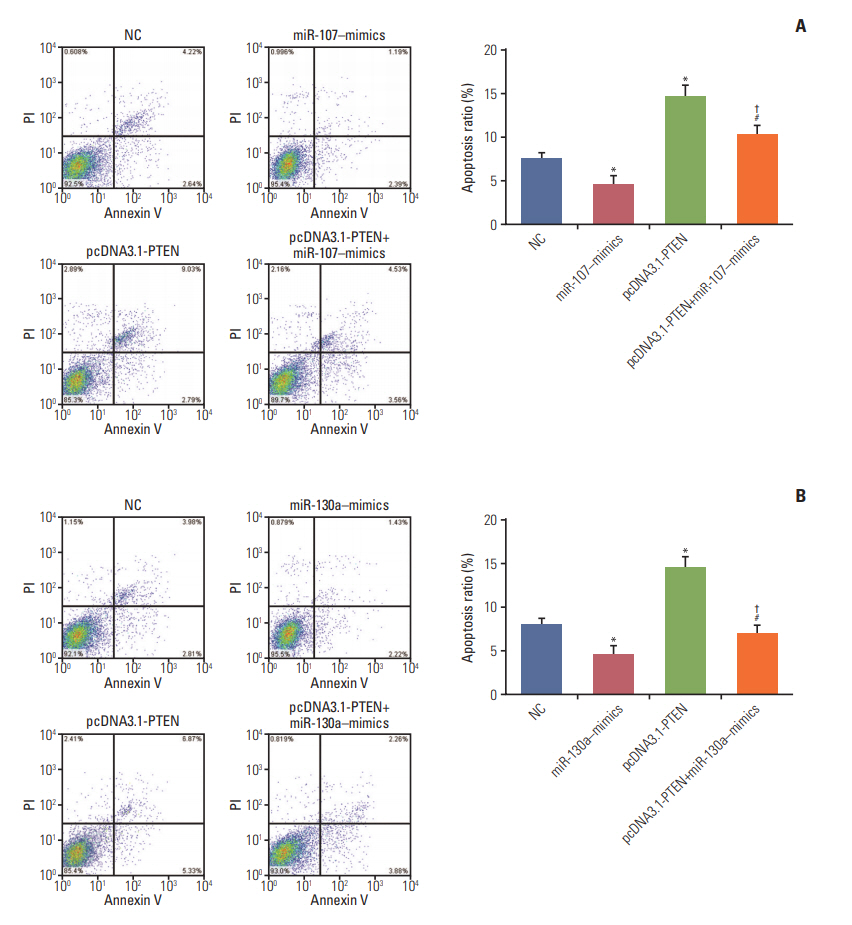
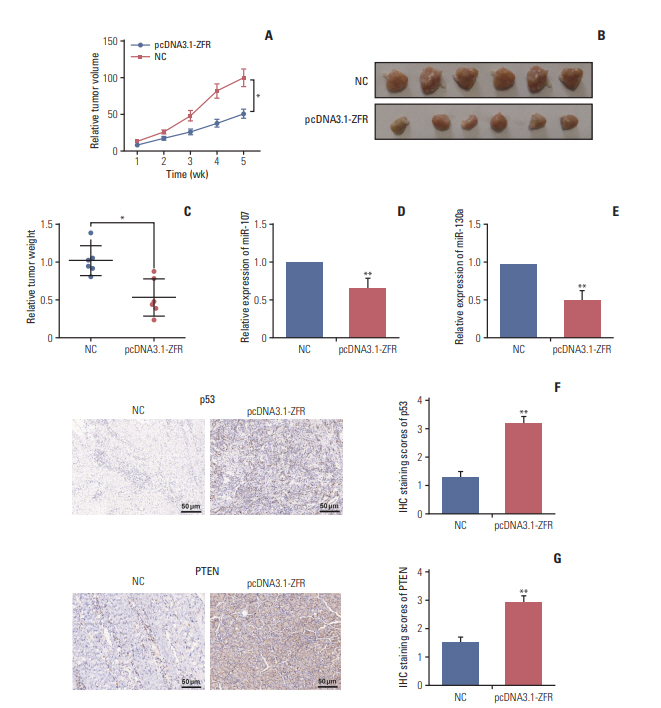
Circ-ZFR and PTEN were low-expressed whereas miR-107 and miR-130a were highexpressed in GC tissues and cells. There existed targeted relationships and interactions between miR-130a/miR-107 and ZFR/PTEN. Circ-ZFR inhibited GC cell propagation, cell cycle and promoted apoptosis by sponging miR-107/miR-130a, while miR-107/miR-130a promoted GC cell propagation and impeded apoptosis through targeting PTEN. Circ-ZFR inhibited cell proliferation and facilitated apoptosis in GC by sponging miR-130a/miR-107 and modulating PTEN. Circ-ZFR curbed GC tumor growth and affected p53 protein expression in vivo.
CONCLUSION
Circ-ZFR restrained GC cell proliferation, induced cell cycle arrest and promoted apoptosis by sponging miR-130a/miR-107 and regulating PTEN.
Keyword
MeSH Terms
Figure
Reference
-
References
1. Xi HQ, Zhang KC, Li JY, Cui JX, Gao YH, Wei B, et al. RNAimediated inhibition of Lgr5 leads to decreased angiogenesis in gastric cancer. Oncotarget. 2017; 8:31581–91.
Article2. Li X, Li H, Zhang R, Liu J, Liu J. MicroRNA-449a inhibits proliferation and induces apoptosis by directly repressing E2F3 in gastric cancer. Cell Physiol Biochem. 2015; 35:2033–42.
Article3. Allemani C, Weir HK, Carreira H, Harewood R, Spika D, Wang XS, et al. Global surveillance of cancer survival 1995-2009: analysis of individual data for 25,676,887 patients from 279 population-based registries in 67 countries (CONCORD-2). Lancet. 2015; 385:977–1010.
Article4. Liu J, Liu T, Wang X, He A. Circles reshaping the RNA world: from waste to treasure. Mol Cancer. 2017; 16:58.
Article5. Chen LL, Yang L. Regulation of circRNA biogenesis. RNA Biol. 2015; 12:381–8.
Article6. Fu L, Wu S, Yao T, Chen Q, Xie Y, Ying S, et al. Decreased expression of hsa_circ_0003570 in hepatocellular carcinoma and its clinical significance. J Clin Lab Anal. 2017 May 11 [Epub]. https://doi.org/10.1002/jcla.22239.
Article7. Huang M, He YR, Liang LC, Huang Q, Zhu ZQ. Circular RNA hsa_circ_0000745 may serve as a diagnostic marker for gastric cancer. World J Gastroenterol. 2017; 23:6330–8.
Article8. Guo JN, Li J, Zhu CL, Feng WT, Shao JX, Wan L, et al. Comprehensive profile of differentially expressed circular RNAs reveals that hsa_circ_0000069 is upregulated and promotes cell proliferation, migration, and invasion in colorectal cancer. Onco Targets Ther. 2016; 9:7451–8.
Article9. Zhang H, Wang G, Ding C, Liu P, Wang R, Ding W, et al. Increased circular RNA UBAP2 acts as a sponge of miR-143 to promote osteosarcoma progression. Oncotarget. 2017; 8:61687–97.
Article10. Izaurralde E. GENE REGULATION. Breakers and blockersmiRNAs at work. Science. 2015; 349:380–2.11. Gattolliat CH, Uguen A, Pesson M, Trillet K, Simon B, Doucet L, et al. MicroRNA and targeted mRNA expression profiling analysis in human colorectal adenomas and adenocarcinomas. Eur J Cancer. 2015; 51:409–20.
Article12. Hansen TB, Kjems J, Damgaard CK. Circular RNA and miR-7 in cancer. Cancer Res. 2013; 73:5609–12.
Article13. Han D, Li J, Wang H, Su X, Hou J, Gu Y, et al. Circular RNA circMTO1 acts as the sponge of microRNA-9 to suppress hepatocellular carcinoma progression. Hepatology. 2017; 66:1151–64.
Article14. Li H, Yang J, Wei X, Song C, Dong D, Huang Y, et al. Circ-FUT10 reduces proliferation and facilitates differentiation of myoblasts by sponging miR-133a. J Cell Physiol. 2018; 233:4643–51.
Article15. Yao JT, Zhao SH, Liu QP, Lv MQ, Zhou DX, Liao ZJ, et al. Over-expression of CircRNA_100876 in non-small cell lung cancer and its prognostic value. Pathol Res Pract. 2017; 213:453–6.
Article16. Fu L, Yao T, Chen Q, Mo X, Hu Y, Guo J. Screening differential circular RNA expression profiles reveals hsa_circ_0004018 is associated with hepatocellular carcinoma. Oncotarget. 2017; 8:58405–16.
Article17. Li P, Chen H, Chen S, Mo X, Li T, Xiao B, et al. Circular RNA 0000096 affects cell growth and migration in gastric cancer. Br J Cancer. 2017; 116:626–33.
Article18. Li J, Yang J, Zhou P, Le Y, Zhou C, Wang S, et al. Circular RNAs in cancer: novel insights into origins, properties, functions and implications. Am J Cancer Res. 2015; 5:472–80.19. Liang HF, Zhang XZ, Liu BG, Jia GT, Li WL. Circular RNA circ-ABCB10 promotes breast cancer proliferation and progression through sponging miR-1271. Am J Cancer Res. 2017; 7:1566–76.20. Tang YY, Zhao P, Zou TN, Duan JJ, Zhi R, Yang SY, et al. Circular RNA hsa_circ_0001982 promotes breast cancer cell carcinogenesis through decreasing miR-143. DNA Cell Biol. 2017; 36:901–8.
Article21. Zhong Z, Lv M, Chen J. Screening differential circular RNA expression profiles reveals the regulatory role of circTCF25-miR-103a-3p/miR-107-CDK6 pathway in bladder carcinoma. Sci Rep. 2016; 6:30919.
Article22. Zou CD, Zhao WM, Wang XN, Li Q, Huang H, Cheng WP, et al. MicroRNA-107: a novel promoter of tumor progression that targets the CPEB3/EGFR axis in human hepatocellular carcinoma. Oncotarget. 2016; 7:266–78.
Article23. Wang S, Ma G, Zhu H, Lv C, Chu H, Tong N, et al. miR-107 regulates tumor progression by targeting NF1 in gastric cancer. Sci Rep. 2016; 6:36531.
Article24. Milella M, Falcone I, Conciatori F, Cesta Incani U, Del Curatolo A, Inzerilli N, et al. PTEN: multiple functions in human malignant tumors. Front Oncol. 2015; 5:24.25. Yu G, Chen X, Chen S, Ye W, Hou K, Liang M. MiR-19a, miR-122 and miR-223 are differentially regulated by hepatitis B virus X protein and involve in cell proliferation in hepatoma cells. J Transl Med. 2016; 14:122.
Article26. Wei H, Cui R, Bahr J, Zanesi N, Luo Z, Meng W, et al. miR-130a deregulates PTEN and stimulates tumor growth. Cancer Res. 2017; 77:6168–78.27. Wang S, Yuan L. Predictive biomarkers for targeted and cytotoxic agents in gastric cancer for personalized medicine. Biosci Trends. 2016; 10:171–80.
Article28. Chen J, Cui L, Yuan J, Zhang Y, Sang H. Circular RNA WDR77 target FGF-2 to regulate vascular smooth muscle cells proliferation and migration by sponging miR-124. Biochem Biophys Res Commun. 2017; 494:126–32.
Article29. Peng L, Chen G, Zhu Z, Shen Z, Du C, Zang R, et al. Circular RNA ZNF609 functions as a competitive endogenous RNA to regulate AKT3 expression by sponging miR-150-5p in Hirschsprung's disease. Oncotarget. 2017; 8:808–18.
Article30. Xie H, Ren X, Xin S, Lan X, Lu G, Lin Y, et al. Emerging roles of circRNA_001569 targeting miR-145 in the proliferation and invasion of colorectal cancer. Oncotarget. 2016; 7:26680–91.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- RETRACTION: Circular RNA-ZFR Inhibited Cell Proliferation and Promoted Apoptosis in Gastric Cancer by Sponging miR-130a/miR-107 and Modulating PTEN
- The Circular RNA Circ_0043947 Promoted Gastric Cancer Progression by Sponging miR-384 to Regulate CREB1 Expression
- Long non-coding RNA T-cell leukemia/lymphoma 6 serves as a sponge for miR-21 modulating the cell proliferation of retinoblastoma through PTEN
- Knockdown of LncRNA H19 Relieves LPS-Induced Damage by Modulating miR-130a in Osteoarthritis
- MSCs-Derived miR-150-5p-Expressing Exosomes Promote Skin Wound Healing by Activating PI3K/AKT Pathway through PTEN