J Gastric Cancer.
2019 Sep;19(3):315-328. 10.5230/jgc.2019.19.e28.
Combined Detection of Serum MiR-221-3p and MiR-122-5p Expression in Diagnosis and Prognosis of Gastric Cancer
- Affiliations
-
- 1Department of Gastroenterology, Taizhou First People's Hospital, Taizhou 318020, China.
- 2Department of General Surgery, Taizhou First People's Hospital, Taizhou 318020, China. liaonansheng2019@163.com
- KMID: 2458831
- DOI: http://doi.org/10.5230/jgc.2019.19.e28
Abstract
- PURPOSE
To investigate the clinical value of serum miR-221-3p and miR-122-5p expression levels in the diagnosis and prognosis of gastric cancer.
MATERIALS AND METHODS
Serum samples from 141 gastric cancer cases (gastric cancer group), 110 gastric polyps (gastric polyp group), and 75 healthy people (healthy control) were used to detect miR-221-3p and miR-122-5p expression using real-time reverse transcription polymerase chain reaction.
RESULTS
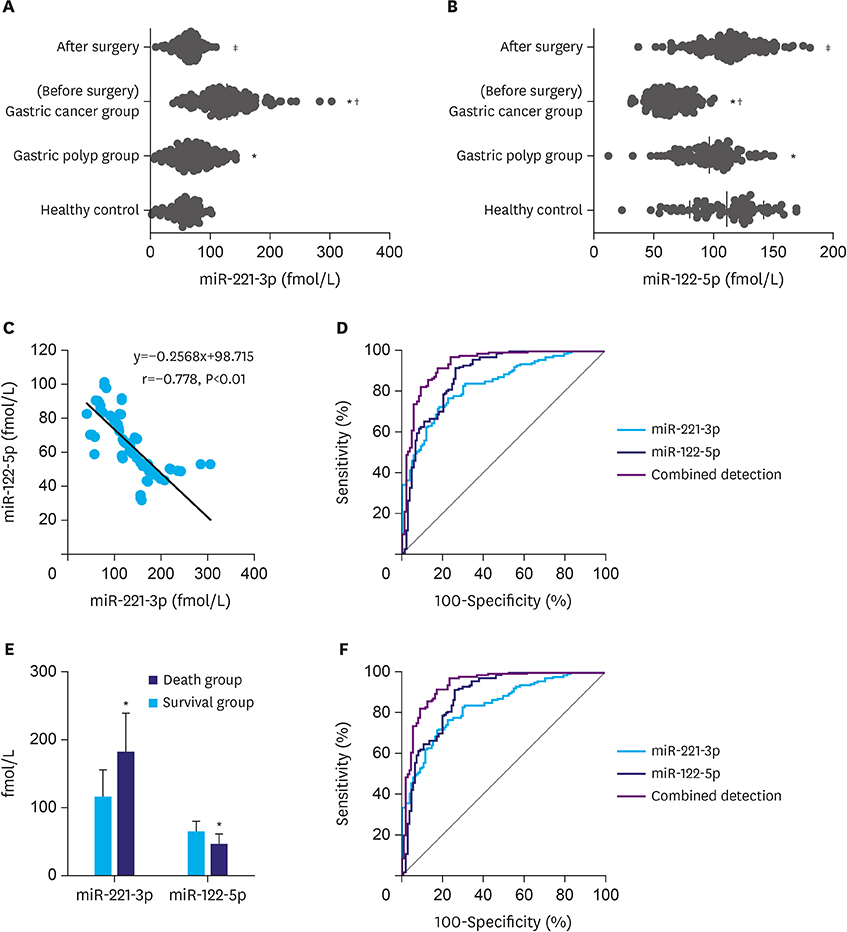
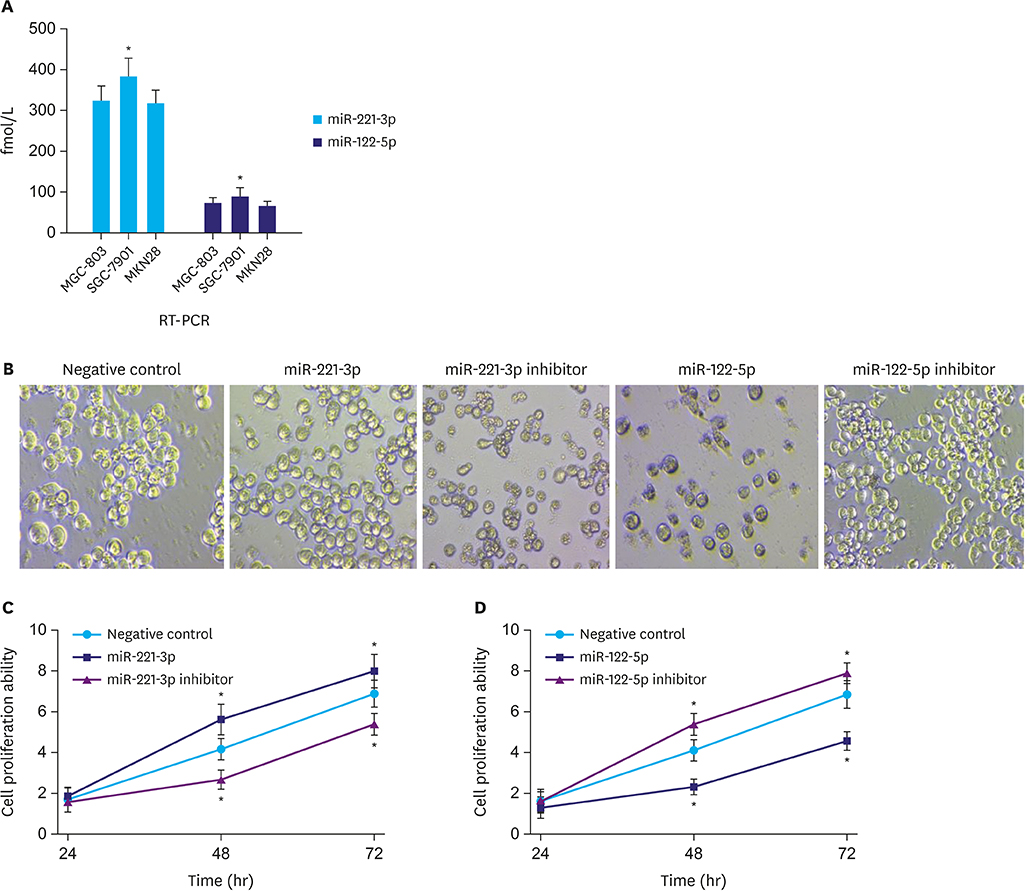
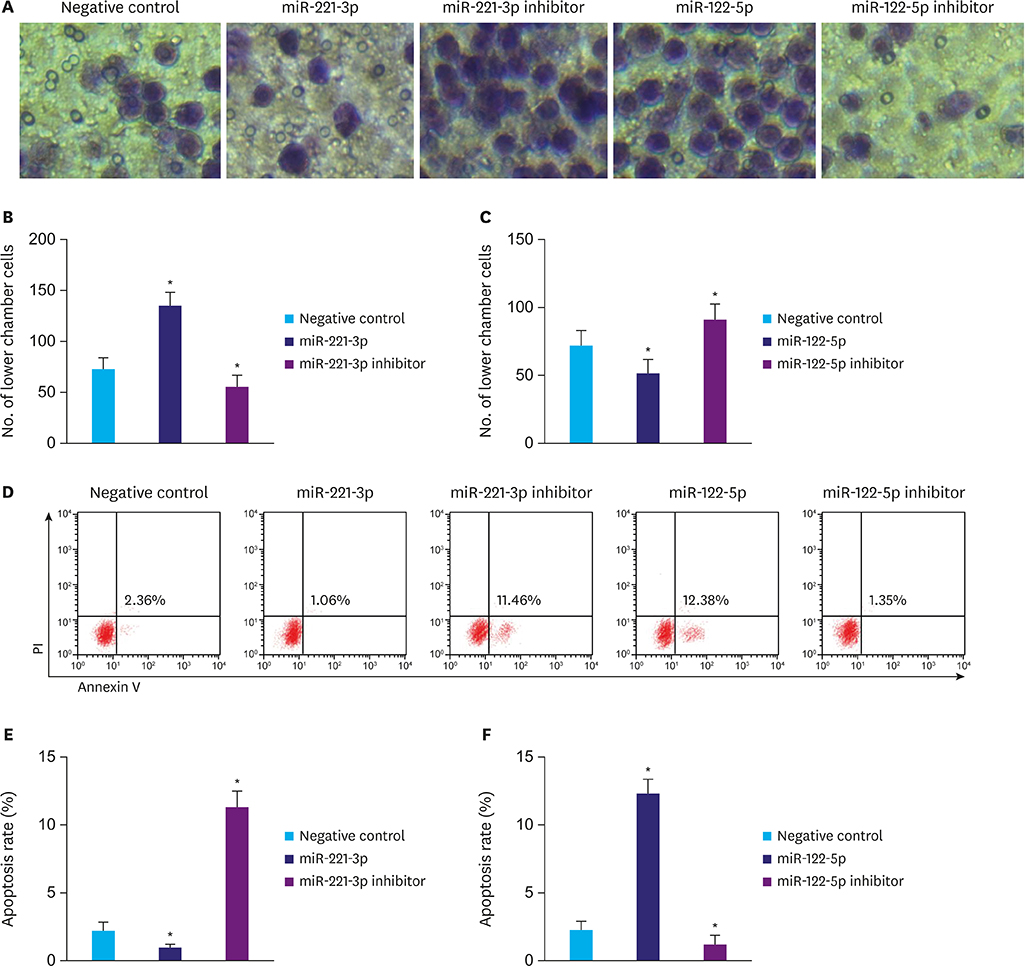
Serum miR-221-3p expression was significantly higher in the gastric cancer group than in the gastric polyp group, and it was significantly lower than that before operation. The miR-221-3p expression was significantly higher in the death group than in the survival group. The proliferation and migration ability significantly increased and the apoptosis rate significantly decreased by miR-221-3p transfection in gastric cancer cells. In contrast, the function of miR-122-5p in gastric cancer cells was opposite of miR-221-3p. Serum miR-221-3p expression was negatively correlated with that of miR-122-5p in gastric cancer. Serum miR-221-3p and miR-122-5p expressions were significantly correlated with the degree of differentiation, tumor, node, metastasis stage, lymph node metastasis, and invasion depth. miR-221-3p and miR-122-5p expression levels were independent prognostic factors for postoperative gastric cancer. In the diagnosis and predicting prognosis of gastric cancer, receiver operating characteristic analysis revealed that the area under curve of combined detection of serum miR-221-3p and miR-122-5p expression had a greater diagnostic effect than either single maker.
CONCLUSIONS
The miR-221-3p and miR-122-5p are involved in the development of gastric cancer, and they have important clinical values in gastric cancer diagnosis and prognosis.
MeSH Terms
Figure
Reference
-
1. Torre LA, Bray F, Siegel RL, Ferlay J, Lortet-Tieulent J, Jemal A. Global cancer statistics, 2012. CA Cancer J Clin. 2015; 65:87–108.
Article2. Gatta G, Rossi S, Capocaccia R. Cancer burden estimates and forecasts: uses and cautions. Tumori. 2013; 99:439–443.
Article3. Fu DG. Epigenetic alterations in gastric cancer (review). Mol Med Rep. 2015; 12:3223–3230.
Article4. Khatami F, Karbakhsh M. Socioeconomic position and incidence of gastric cancer: a systematic review and meta-analysis. J Epidemiol Community Health. 2015; 69:818–819.
Article5. Yoon H, Kim N. Diagnosis and management of high risk group for gastric cancer. Gut Liver. 2015; 9:5–17.
Article6. Stojanovic J, Tognetto A, Tiziano DF, Leoncini E, Posteraro B, Pastorino R, et al. MicroRNAs expression profiles as diagnostic biomarkers of gastric cancer: a systematic literature review. Biomarkers. 2019; 24:110–119.
Article7. Yuan HL, Wang T, Zhang KH. MicroRNAs as potential biomarkers for diagnosis, therapy and prognosis of gastric cancer. Onco Targets Ther. 2018; 11:3891–3900.
Article8. Wu Q, Ren X, Zhang Y, Fu X, Li Y, Peng Y, et al. MiR-221-3p targets ARF4 and inhibits the proliferation and migration of epithelial ovarian cancer cells. Biochem Biophys Res Commun. 2018; 497:1162–1170.
Article9. Deng L, Lei Q, Wang Y, Wang Z, Xie G, Zhong X, et al. Downregulation of miR-221-3p and upregulation of its target gene PARP1 are prognostic biomarkers for triple negative breast cancer patients and associated with poor prognosis. Oncotarget. 2017; 8:108712–108725.
Article10. Tao K, Yang J, Guo Z, Hu Y, Sheng H, Gao H, et al. Prognostic value of miR-221-3p, miR-342-3p and miR-491-5p expression in colon cancer. Am J Transl Res. 2014; 6:391–401.11. Xu X, Gao F, Wang J, Tao L, Ye J, Ding L, et al. MiR-122-5p inhibits cell migration and invasion in gastric cancer by down-regulating DUSP4. Cancer Biol Ther. 2018; 19:427–435.
Article12. Barajas JM, Reyes R, Guerrero MJ, Jacob ST, Motiwala T, Ghoshal K. The role of miR-122 in the dysregulation of glucose-6-phosphate dehydrogenase (G6PD) expression in hepatocellular cancer. Sci Rep. 2018; 8:9105.
Article13. Duan Y, Dong Y, Dang R, Hu Z, Yang Y, Hu Y, et al. MiR-122 inhibits epithelial mesenchymal transition by regulating P4HA1 in ovarian cancer cells. Cell Biol Int. 2018; 42:1564–1574.
Article14. Ergün S, Ulasli M, Igci YZ, Igci M, Kırkbes S, Borazan E, et al. The association of the expression of miR-122-5p and its target ADAM10 with human breast cancer. Mol Biol Rep. 2015; 42:497–505.
Article15. Heinemann FG, Tolkach Y, Deng M, Schmidt D, Perner S, Kristiansen G, et al. Serum miR-122-5p and miR-206 expression: non-invasive prognostic biomarkers for renal cell carcinoma. Clin Epigenetics. 2018; 10:11.
Article16. Uen Y, Wang JW, Wang C, Jhang Y, Chung JY, Tseng T, et al. Mining of potential microRNAs with clinical correlation - regulation of syndecan-1 expression by miR-122-5p altered mobility of breast cancer cells and possible correlation with liver injury. Oncotarget. 2018; 9:28165–28175.
Article17. Pei ZJ, Zhang ZG, Hu AX, Yang F, Gai Y. miR-122-5p inhibits tumor cell proliferation and induces apoptosis by targeting MYC in gastric cancer cells. Pharmazie. 2017; 72:344–347.18. Mitchell PS, Parkin RK, Kroh EM, Fritz BR, Wyman SK, Pogosova-Agadjanyan EL, et al. Circulating microRNAs as stable blood-based markers for cancer detection. Proc Natl Acad Sci U S A. 2008; 105:10513–10518.
Article19. Clancy C, Joyce MR, Kerin MJ. The use of circulating microRNAs as diagnostic biomarkers in colorectal cancer. Cancer Biomark. 2015; 15:103–113.
Article20. Li M, Zhou Y, Xia T, Zhou X, Huang Z, Zhang H, et al. Circulating microRNAs from the miR-106a-363 cluster on chromosome X as novel diagnostic biomarkers for breast cancer. Breast Cancer Res Treat. 2018; 170:257–270.
Article21. Wang X, Jia Z, Shi H, Pan C. Identification and evaluation of 2 circulating microRNAs for non-small cell lung cancer diagnosis. Clin Exp Pharmacol Physiol. 2018; 45:1083–1086.
Article22. Ho AS, Huang X, Cao H, Christman-Skieller C, Bennewith K, Le QT, et al. Circulating miR-210 as a novel hypoxia marker in pancreatic cancer. Transl Oncol. 2010; 3:109–113.
Article23. Jones LB, Bell CR, Bibb KE, Gu L, Coats MT, Matthews QL. Pathogens and their effect on exosome biogenesis and composition. Biomedicines. 2018; 6:E79.
Article24. Kosaka N, Iguchi H, Yoshioka Y, Takeshita F, Matsuki Y, Ochiya T. Secretory mechanisms and intercellular transfer of microRNAs in living cells. J Biol Chem. 2010; 285:17442–17452.
Article25. Wu XG, Zhou CF, Zhang YM, Yan RM, Wei WF, Chen XJ, et al. Cancer-derived exosomal miR-221-3p promotes angiogenesis by targeting THBS2 in cervical squamous cell carcinoma. Angiogenesis. 2019; 22:397–410.
Article26. Zhou CF, Ma J, Huang L, Yi HY, Zhang YM, Wu XG, et al. Cervical squamous cell carcinoma-secreted exosomal miR-221-3p promotes lymphangiogenesis and lymphatic metastasis by targeting VASH1. Oncogene. 2019; 38:1256–1268.
Article27. Liu YH, Liu JL, Wang Z, Zhu XH, Chen XB, Wang MQ. MiR-122-5p suppresses cell proliferation, migration and invasion by targeting SATB1 in nasopharyngeal carcinoma. Eur Rev Med Pharmacol Sci. 2019; 23:622–629.28. Xu Z, Liu G, Zhang M, Zhang Z, Jia Y, Peng L, et al. miR-122-5p inhibits the proliferation, invasion and growth of bile duct carcinoma cells by targeting ALDOA. Cell Physiol Biochem. 2018; 48:2596–2606.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Expression of specific microRNAs in tissue and plasma in colorectal cancer
- Serum miR-329-3p as a potential biomarker for poor ovarian response in an in vitro fertilization
- miR-132 and miR-942 Expression Levels in Children with Attention Deficit and Hyperactivity Disorder: A Controlled Study
- LncRNA XLOC_006390 facilitates cervical cancer tumorigenesis and metastasis as a ceRNA against miR-331-3p and miR-338-3p
- Expression of miR-221 and miR-18a in patients with hepatocellular carcinoma and its clinical significance