Tuberc Respir Dis.
2007 Jul;63(1):52-58. 10.4046/trd.2007.63.1.52.
Gene Expression Profile of Lung Cancer Cells Following Photodynamic Therapy
- Affiliations
-
- 1Department of Molecular and Cellular Biochemistry, College of Medicine, Kangwon National University, Korea.
- 2Clinical Research Institute of Kangwon National University Hospital, Korea.
- 3Department of Internal Medicine, College of Medicine, Kangwon National University, Chunchon, Korea. pulmo2@kangwon.ac.kr
- KMID: 1877317
- DOI: http://doi.org/10.4046/trd.2007.63.1.52
Abstract
-
BACKGROUND: Photodynamic therapy is a viable option for lung cancer treatment, and many studies have shown that it is capable of inducing cell death in lung cancer cells. However, the precise mechanism of this cell death has not been fully elucidated. To investigate the early changes in cancer cell transcription, we treated A549 cells with the photosensitizer DH-I-180-3 and then we illuminated the cells.
METHODS
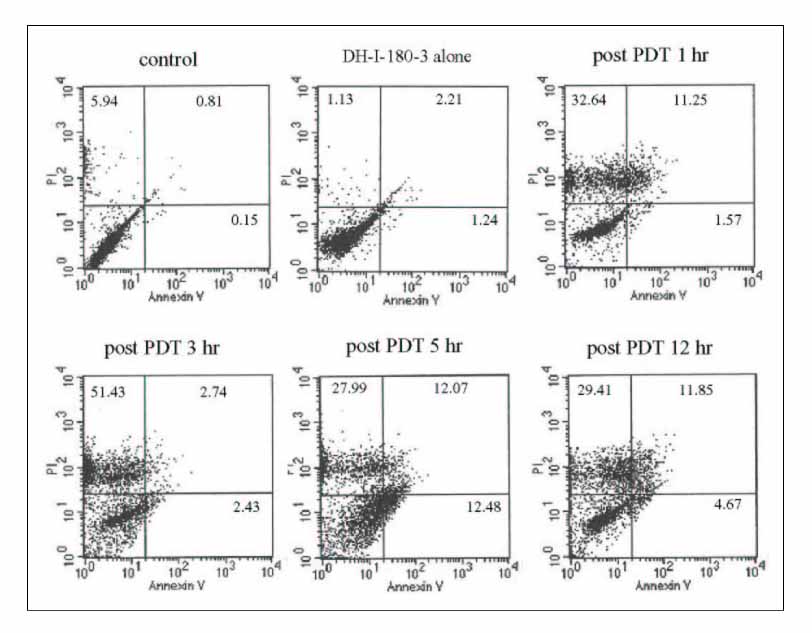
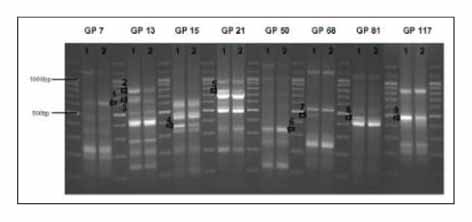
We investigated the gene expression profiles of the the A549 lung cancer cell line, using a DEG kit, following photodynamic therapy and we evaluated the cell viability by performing flow cytometry. We identified the genes that were significantly changed following photodynamic therapy by performing DNA sequencing.
RESULTS
The FACS data showed that the cell death of the lung cancer cells was mainly caused by necrosis. We found nine genes that were significantly changed and we identified eight of these genes. We evaluated the expression of two genes, 3-phosphoglycerate dehydrogenase and ribosomal protein S29. The expressed level of carbonic anhydrase XII, clusterin, MRP3s1 protein, complement 3, membrane cofactor protein and integrin beta 1 were decreased.
CONCLUSION
Many of the gene products are membrane-associated proteins. The main mechanism of photodynamic therapy with using the photosensitizing agent DH-I-180-3 appears to be necrosis and this may be associated with the altered production of membrane proteins.
MeSH Terms
-
Antigens, CD46
Carbonic Anhydrases
Cell Death
Cell Line
Cell Survival
Clusterin
Complement System Proteins
Flow Cytometry
Gene Expression Profiling
Gene Expression*
Lung Neoplasms*
Lung*
Membrane Proteins
Necrosis
Phosphoglycerate Dehydrogenase
Photochemotherapy*
Photosensitizing Agents
Ribosomal Proteins
Sequence Analysis, DNA
Transcriptome*
Antigens, CD46
Carbonic Anhydrases
Clusterin
Complement System Proteins
Membrane Proteins
Phosphoglycerate Dehydrogenase
Photosensitizing Agents
Ribosomal Proteins
Figure
Reference
-
1. Spiro SG, Porter JC. Lung cancer--where are we today? Current advances in staging and nonsurgical treatment. Am J Respir Crit Care Med. 2002. 166:1166–1196.2. Spira A, Ettinger DS. Multidisciplinary management of lung cancer. N Engl J Med. 2004. 350:379–392.3. Moghissi K, Dixon K. Is bronchoscopic photodynamic therapy a therapeutic option in lung cancer? Eur Respir J. 2003. 22:535–541.4. Dougherty TJ, Gomer CJ, Henderson BW, Jori G, Kessel D, Korbelik M, et al. Photodynamic therapy. J Natl Cancer Inst. 1998. 90:889–905.5. Kim CS, Lee CH, Lee PH, Han S. Inactivation of mitochondrial electron transport by photosensitization of a pheophorbide a derivative. Mol Cells. 2004. 17:347–352.6. Lim DS, Ko SH, Won DH, Lee CH, Lee WY. Photodynamic anti-tumor activity of a new chlorine-based photosensitizer against Lewis lung carcinoma cells in vitro and in vivo. J Porphyrins Phthalocyanines. 2003. 7:155–161.7. Kim YJ, Kwak CI, Gu YY, Hwang IT, Chun JY. Annealing control primer system for identification of differentially expressed genes on agarose gels. Biotechniques. 2004. 36:424–430.8. Cui XS, Shin MR, Lee KA, Kim NH. Identification of differentially expressed genes in murine embryos at the blastocyst stage using annealing control primer system. Mol Reprod Dev. 2005. 70:278–287.9. Fabris C, Valduga G, Miotto G, Borsetto L, Jori G, Garbisa S, et al. Photosensitization with zinc (II) phthalocyanine as a switch in the decision between apoptosis and necrosis. Cancer Res. 2001. 61:7495–7500.10. Dellinger M. Apoptosis or necrosis following photofrin photosensitization: influence of the incubation protocol. Photochem Photobiol. 1996. 64:182–187.11. Vermes I, Haanen C, Steffens-Nakken H, Reutelingsperger C. A novel assay for apoptosis. Flow cytometric detection of phosphatidylserine expression on early apoptotic cells using fluorescein labeled Annexin V. J Immunol Methods. 1995. 184:39–51.12. Ahn WS, Bae SM, Huh SW, Lee JM, Namkoong SE, Han SJ, et al. Necrosis-like death with plasma membrane damage against cervical cancer cells by photodynamic therapy. Int J Gynecol Cancer. 2004. 14:475–482.13. Ulmasov B, Waheed A, Shah GN, Grubb JH, Sly WS, Tu C, et al. Purification and kinetic analysis of recombinant CA XII, a membrane carbonic anhydrase overexpressed in certain cancers. Proc Natl Acad Sci USA. 2000. 97:14212–14217.14. Leppilampi M, Saarnio J, Karttunen TJ, Kivela J, Pastorekova S, Pastorek J, et al. Carbonic anhydrase isoenzymes IX and XII in gastric tumors. World J Gastroenterol. 2003. 9:1398–1403.15. Supuran CT, Scozzafava A, Casini A. Carbonic anhydrase inhibitors. Med Res Rev. 2003. 23:146–189.16. Torky AR, Stehfest E, Viehweger K, Taege C, Foth H. Immuno-histochemical detection of MRPs in human lung cells in culture. Toxicology. 2005. 207:437–450.17. Young LC, Campling BG, Cole SP, Deeley RG, Gerlach JH. Multidrug resistance proteins MRP3, MRP1, and MRP2 in lung cancer: correlation of protein levels with drug response and messenger RNA levels. Clin Cancer Res. 2001. 7:1798–1804.18. Oshita F, Kameda Y, Ikehara M, Tanaka G, Yamada K, Nomura I, et al. Increased expression of integrin beta 1 is poor prognostic factor in small-cell lung cancer. Anticancer Res. 2002. 22:1065–1070.19. Cordes N, Blaese MA, Meineke V, Van Beuningen D. Ionizing radiation induces up-regulation of functional beta1-integrin in human lung tumour cell lines in vitro. Int J Radiat Biol. 2002. 78:347–357.20. Park CC, Zhang H, Pallavicini M, Gray JW, Baehner F, Park CJ, et al. Beta1 integrin inhibitory antibody induced apoptosis of breast cancer cells, inhibits growth, and distinguished malignant from normal phenotype in three dimensional cultures and in vivo. Cancer Res. 2006. 66:1526–1535.21. Hakulinen J, Junnikkala S, Sorsa T, Meri S. Complement inhibitor membrane cofactor protein is constitutively shed from cancer cell membrane in vesicles and converted by a metalloproteinase to a functionally active soluble form. Eur J Immunol. 2004. 34:2620–2629.22. Madjd Z, Durrant LG, Pinder SE, Ellis IO, Ronan J, Lewis S, et al. Do poor-prognosis breast tumours express membrane cofactor proteins (CD46)? Cancer Immunol Immunother. 2005. 54:149–156.23. Cecic I, Serrano K, Gyongyossy-Issa M, Korbelik M. Characteristics of complement activation in mice bearing Lewis lung carcinomas treated by photodynamic therapy. Cancer Lett. 2005. 225:215–223.24. Shannan B, Seifert M, Leskov K, Willis J, Boothman D, Tilgen W, et al. Challenge and promise: roles for clusterin in pathogenesis, progression and therapy of cancer. Cell Death Differ. 2006. 13:12–19.25. Pajak B, Orzechowski A. Clusterin: the missing link in the calcium-dependent resistance of cancer cells to apoptogenic stimuli. Postepy Hig Med Dosw. 2006. 60:45–51.26. July LV, Beraldi E, So A, Fazli L, Evans K, English JC, et al. Nucleotide-based therapies targeting clusterin chemosensitize human lung adenocarcinoma cells both in vitro and in vivo. Mol Cancer Ther. 2004. 3:223–232.27. Cho HM, Jun DY, Bae MA, Ahn JD, Kim YH. Nucleotide sequence and differential expression of the human 3-phosphoglycerate dehydrogenase gene. Gene. 2000. 245:193–201.28. Khanna N, Reddy VG, Tuteja N, Singh N. Differential gene expression in apoptosis: identification of ribosomal protein S29 as an apoptotic inducer. Biochem Biophys Res Commun. 2000. 277:476–486.29. Zhou ZD, Bao L, Liu DG, Li MQ, Ge YZ, Huang YL, et al. Low content of protein S29 in ribosomes of human lung cancer cell line a549: detected by two-dimensional electrophoresis. Protein Pept Lett. 2003. 10:91–97.30. Khanna N, Sen S, Sharma H, Singh N. S29 ribosomal protein induces apoptosis in H520 cells and sensitizes them to chemotherapy. Biochem Biophys Res Commun. 2003. 304:26–35.31. Wild PJ, Krieg RC, Seidl J, Stoehr R, Reher K, Hofmann C, et al. RNA expression profiling of normal and tumor cells following photodynamic therapy with 5-aminolevulinic acid-induced protoporphyrin IX in vitro. Mol Cancer Ther. 2005. 4:516–528.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Usefulness of Photodynamic Therapy in the Management of Early Central Lung Cancer: A Report of Three Cases
- Use of a Combined Gene Expression Profile in Implementing a Drug Sensitivity Predictive Model for Breast Cancer
- Photodynamic Therapy in Early Lung Cancer: A Report of Two Cases
- Study on the Function of NAG-1 in Hepatocellular and Gastric Carcinoma Cells
- Photodynamic Therapy for Endobronchial Obstruction due to Recurrent Lung Cancer : 2 Cases Report