Cancer Res Treat.
2024 Apr;56(2):484-501. 10.4143/crt.2023.712.
Revolutionizing Non–Small Cell Lung Cancer Diagnosis: Ultra-High-Sensitive ctDNA Analysis for Detecting Hotspot Mutations with Long-term Stored Plasma
- Affiliations
-
- 1Department of Medical Science, Asan Medical Institute of Convergence Science and Technology, Seoul, Korea
- 2Department of Pathology, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
- 3Department of Pulmonary and Critical Care Medicine, Asan Medical Center, University of Ulsan College of Medicine, Seoul, Korea
- KMID: 2554338
- DOI: http://doi.org/10.4143/crt.2023.712
Abstract
- Purpose
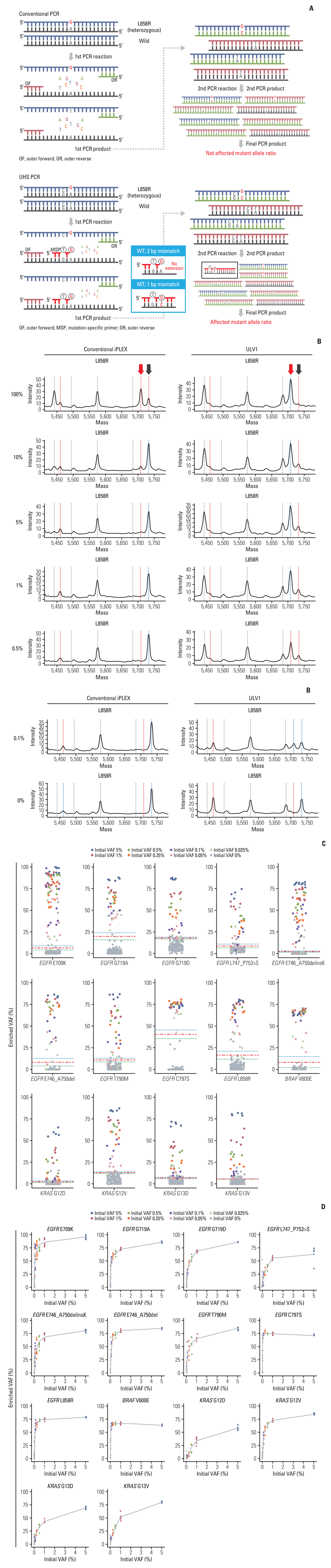
Circulating cell-free DNA (cfDNA) has great potential in clinical oncology. The prognostic and predictive values of cfDNA in non–small cell lung cancer (NSCLC) have been reported, with epidermal growth factor receptor (EGFR), KRAS, and BRAF mutations in tumor-derived cfDNAs acting as biomarkers during the early stages of tumor progression and recurrence. However, extremely low tumor-derived DNA rates hinder cfDNA application. We developed an ultra-high-sensitivity lung version 1 (ULV1) panel targeting BRAF, KRAS, and EGFR hotspot mutations using small amounts of cfDNA, allowing for semi-quantitative analysis with excellent limit-of-detection (0.05%).
Materials and Methods
Mutation analysis was performed on cfDNAs extracted from the plasma of 104 patients with NSCLC by using the ULV1 panel and targeted next-generation sequencing (CT-ULTRA), followed by comparison analysis of mutation patterns previously screened using matched tumor tissue DNA.
Results
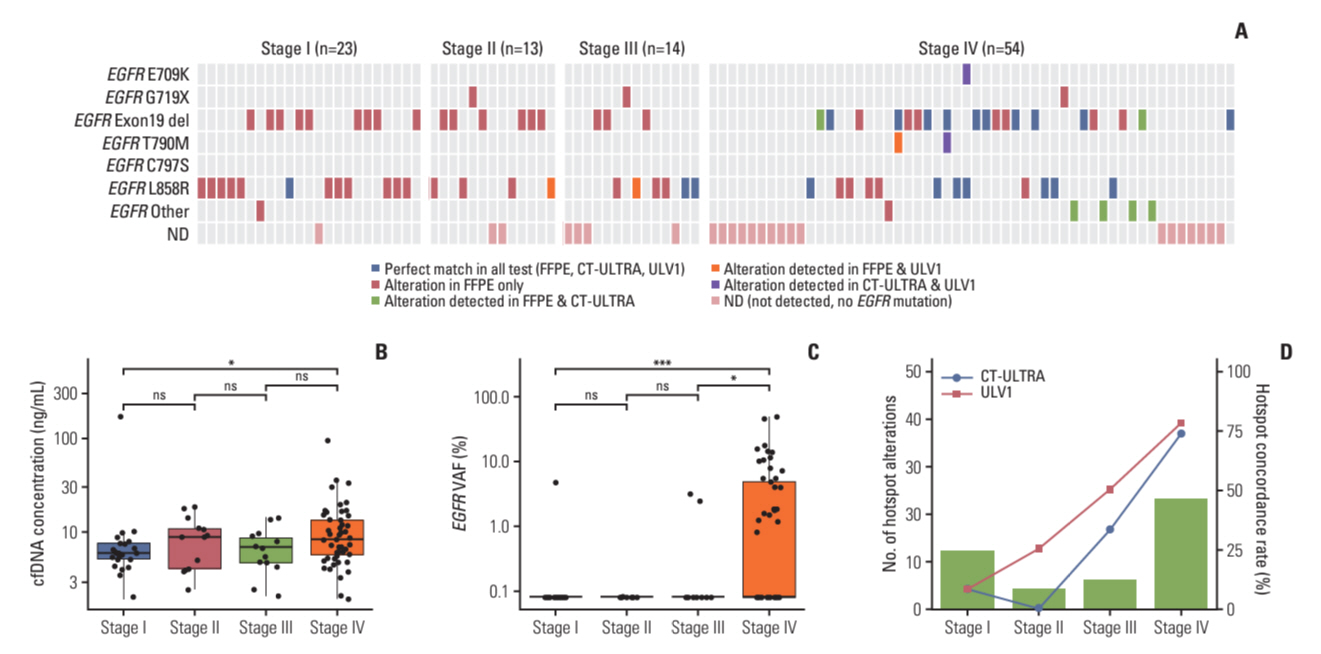
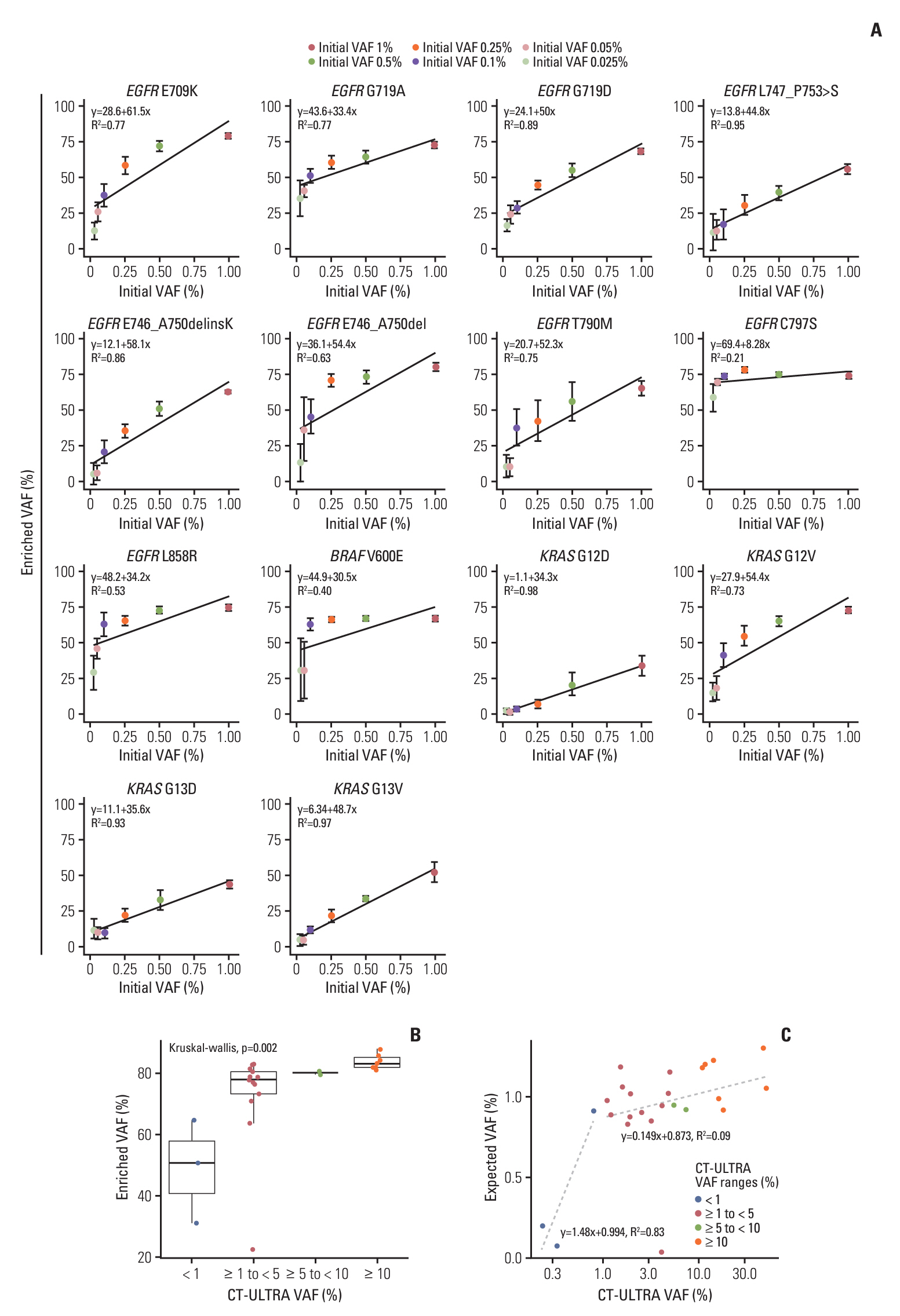
The ULV1 panel demonstrated robust selective amplification of mutant alleles, enabling the detection of mutations with a high degree of analytical sensitivity (limit-of-detection, 0.025%-0.1%) and specificity (87.9%-100%). Applying ULV1 to NSCLC cfDNA revealed 51.1% (23/45) samples with EGFR mutations, increasing with tumor stage: 8.33% (stage I) to 78.26% (stage IV). Semi-quantitative analysis proved effective for low-mutation-fraction clinical samples. Comparative analysis with PANAMutyper EGFR exhibited substantial concordance (κ=0.84).
Conclusion
Good detection sensitivity (~80%) was observed despite the limited volume (1 mL) and long-term storage (12-50 months) of plasma used and is expected to increase with high cfDNA inputs. Thus, the ULV1 panel is a fast and cost-effective method for early diagnosis, treatment selection, and clinical follow-up of patients with NSCLC.
Keyword
Figure
Reference
-
References
1. Lynch TJ, Bell DW, Sordella R, Gurubhagavatula S, Okimoto RA, Brannigan BW, et al. Activating mutations in the epidermal growth factor receptor underlying responsiveness of non-small-cell lung cancer to gefitinib. N Engl J Med. 2004; 350:2129–39.
Article2. Yuan M, Huang LL, Chen JH, Wu J, Xu Q. The emerging treatment landscape of targeted therapy in non-small-cell lung cancer. Signal Transduct Target Ther. 2019; 4:61.
Article3. Wang S, Cang S, Liu D. Third-generation inhibitors targeting EGFR T790M mutation in advanced non-small cell lung cancer. J Hematol Oncol. 2016; 9:34.
Article4. Wang S, Tsui ST, Liu C, Song Y, Liu D. EGFR C797S mutation mediates resistance to third-generation inhibitors in T790M-positive non-small cell lung cancer. J Hematol Oncol. 2016; 9:59.
Article5. Taus A, Camacho L, Rocha P, Hardy-Werbin M, Pijuan L, Piquer G, et al. Dynamics of EGFR mutation load in plasma for prediction of treatment response and disease progression in patients with EGFR-mutant lung adenocarcinoma. Clin Lung Cancer. 2018; 19:387–94.
Article6. Zhang JT, Liu SY, Gao W, Liu SM, Yan HH, Ji L, et al. Longitudinal undetectable molecular residual disease defines potentially cured population in localized non-small cell lung cancer. Cancer Discov. 2022; 12:1690–701.
Article7. Russano M, Napolitano A, Ribelli G, Iuliani M, Simonetti S, Citarella F, et al. Liquid biopsy and tumor heterogeneity in metastatic solid tumors: the potentiality of blood samples. J Exp Clin Cancer Res. 2020; 39:95.
Article8. Bronkhorst AJ, Ungerer V, Holdenrieder S. The emerging role of cell-free DNA as a molecular marker for cancer management. Biomol Detect Quantif. 2019; 17:100087.
Article9. Yi X, Ma J, Guan Y, Chen R, Yang L, Xia X. The feasibility of using mutation detection in ctDNA to assess tumor dynamics. Int J Cancer. 2017; 140:2642–7.
Article10. Corcoran RB, Chabner BA. Application of cell-free DNA analysis to cancer treatment. N Engl J Med. 2018; 379:1754–65.
Article11. Shin SJ, Chun SM, Kim TI, Kim YJ, Choi HJ, Jang SJ, et al. Feasibility of multiplexed gene mutation detection in plasma samples of colorectal cancer patients by mass spectrometric genotyping. PLoS One. 2017; 12:e0176340.
Article12. Elazezy M, Joosse SA. Techniques of using circulating tumor DNA as a liquid biopsy component in cancer management. Comput Struct Biotechnol J. 2018; 16:370–8.
Article13. Sozzi G, Roz L, Conte D, Mariani L, Andriani F, Verderio P, et al. Effects of prolonged storage of whole plasma or isolated plasma DNA on the results of circulating DNA quantification assays. J Natl Cancer Inst. 2005; 97:1848–50.
Article14. Jahangiri L, Hurst T. Assessing the concordance of genomic alterations between circulating-free DNA and tumour tissue in cancer patients. Cancers (Basel). 2019; 11:1938.
Article15. Guo Q, Wang J, Xiao J, Wang L, Hu X, Yu W, et al. Heterogeneous mutation pattern in tumor tissue and circulating tumor DNA warrants parallel NGS panel testing. Mol Cancer. 2018; 17:131.
Article16. Jiang J, Adams HP, Yao L, Yaung S, Lal P, Balasubramanyam A, et al. Concordance of genomic alterations by next-generation sequencing in tumor tissue versus cell-free DNA in stage I-IV non-small cell lung cancer. J Mol Diagn. 2020; 22:228–35.
Article17. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014; 6:224ra24.18. Kujala J, Hartikainen JM, Tengstrom M, Sironen R, Auvinen P, Kosma VM, et al. Circulating cell-free DNA reflects the clonal evolution of breast cancer tumors. Cancers (Basel). 2022; 14:1332.
Article19. Kujala J, Hartikainen JM, Tengstrom M, Sironen R, Kosma VM, Mannermaa A. High mutation burden of circulating cell-free DNA in early-stage breast cancer patients is associated with a poor relapse-free survival. Cancer Med. 2020; 9:5922–31.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Evaluation of Circulating Tumor DNA in Patients with Ovarian Cancer Harboring Somatic PIK3CA or KRAS Mutations
- Analytical and Clinical Validation of a Highly Sensitive NGS-Based ctDNA Assay with Real-World Concordance in Non–Small Cell Lung Cancer
- Feasibility of Circulating Tumor DNA Analysis in Patients with Follicular Lymphoma
- Utilizing Plasma Circulating Tumor DNA Sequencing for Precision Medicine in the Management of Solid Cancers
- Clinical Circulating Tumor DNA Testing for Precision Oncology