Tuberc Respir Dis.
2016 Jul;79(3):165-178. 10.4046/trd.2016.79.3.165.
Lung Microbiome Analysis in Steroid-NaÑ—ve Asthma Patients by Using Whole Sputum
- Affiliations
-
- 1Department of Internal Medicine, Chung-Ang University College of Medicine, Seoul, Korea.
- 2Department of Internal Medicine, Seoul National University College of Medicine, Seoul, Korea. helenmed@snu.ac.kr
- 3Institute of Allergy and Clinical Immunology, Seoul National University Medical Research Center, Seoul, Korea.
- 4Department of Microbiology, Chung-Ang University College of Medicine, Seoul, Korea. kimkj@cau.ac.kr
- KMID: 2326685
- DOI: http://doi.org/10.4046/trd.2016.79.3.165
Abstract
- BACKGROUND
Although recent metagenomic approaches have characterized the distinguished microbial compositions in airways of asthmatics, these results did not reach a consensus due to the small sample size, non-standardization of specimens and medication status. We conducted a metagenomics approach by using terminal restriction fragment length polymorphism (T-RFLP) analysis of the induced whole sputum representing both the cellular and fluid phases in a relative large number of steroid naïve asthmatics.
METHODS
Induced whole sputum samples obtained from 36 healthy subjects and 89 steroid-naÑ—ve asthma patients were analyzed through T-RFLP analysis.
RESULTS
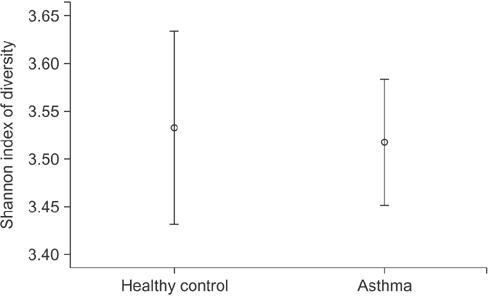
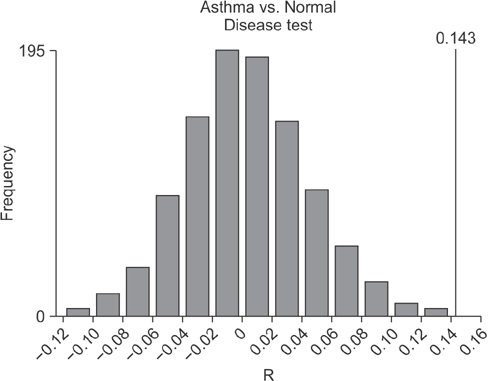
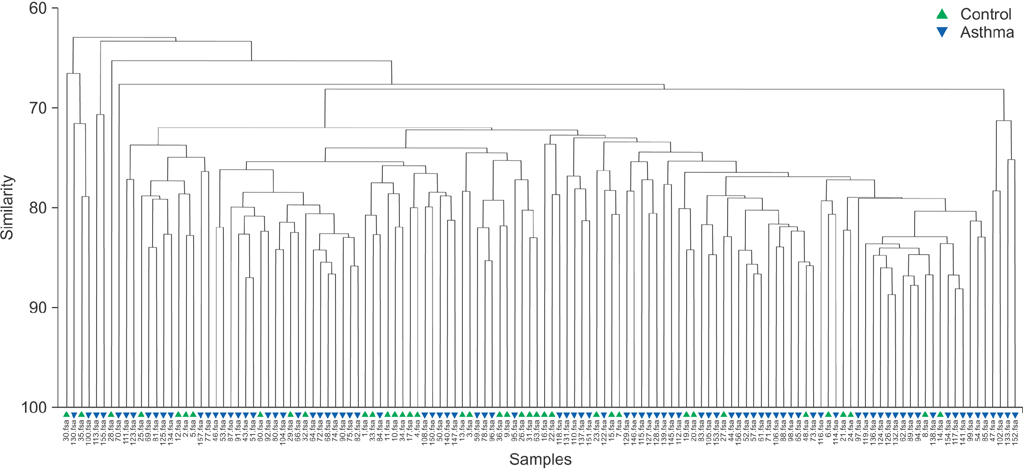
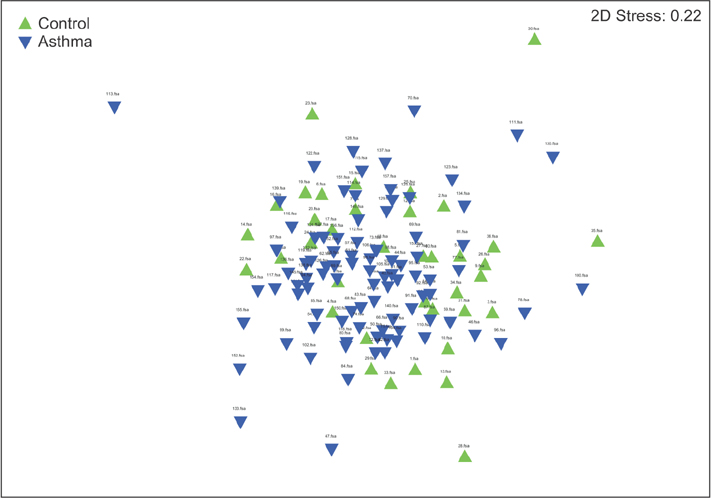
In contrast to previous reports about microbiota in the asthmatic airways, the diversity of microbial composition was not significantly different between the controls and asthma patients (p=0.937). In an analysis of similarities, the global R-value showed a statistically significant difference but a very low separation (0.148, p=0.002). The dissimilarity in the bacterial communities between groups was 28.74%, and operational taxonomic units (OTUs) contributing to this difference were as follows: OTU 789 (Lachnospiraceae), 517 (Comamonadaceae, Acetobacteraceae , and Chloroplast), 633 (Prevotella), 645 (Actinobacteria and Propionibacterium acnes), 607 (Lactobacillus buchneri, Lactobacillus otakiensis, Lactobacillus sunkii, and Rhodobacteraceae), and 661 (Acinetobacter, Pseudomonas, and Leptotrichiaceae), and they were significantly more prevalent in the sputum of asthma patients than in the sputum of the controls.
CONCLUSION
Before starting anti-asthmatic treatment, the microbiota in the whole sputum of patients with asthma showed a marginal difference from the microbiota in the whole sputum of the controls.
MeSH Terms
Figure
Reference
-
1. Wenzel SE. Asthma: defining of the persistent adult phenotypes. Lancet. 2006; 368:804–813.2. Cosentini R, Tarsia P, Canetta C, Graziadei G, Brambilla AM, Aliberti S, et al. Severe asthma exacerbation: role of acute Chlamydophila pneumoniae and Mycoplasma pneumoniae infection. Respir Res. 2008; 9:48.3. Bisgaard H, Hermansen MN, Buchvald F, Loland L, Halkjaer LB, Bonnelykke K, et al. Childhood asthma after bacterial colonization of the airway in neonates. N Engl J Med. 2007; 357:1487–1495.4. Huang YJ, Boushey HA. The microbiome in asthma. J Allergy Clin Immunol. 2015; 135:25–30.5. Holt PG. The mechanism or mechanisms driving atopic asthma initiation: the infant respiratory microbiome moves to center stage. J Allergy Clin Immunol. 2015; 136:15–22.6. Panzer AR, Lynch SV. Influence and effect of the human microbiome in allergy and asthma. Curr Opin Rheumatol. 2015; 27:373–380.7. Edwards MR, Bartlett NW, Hussell T, Openshaw P, Johnston SL. The microbiology of asthma. Nat Rev Microbiol. 2012; 10:459–471.8. Huang YJ. The respiratory microbiome and innate immunity in asthma. Curr Opin Pulm Med. 2015; 21:27–32.9. Althani AA, Marei HE, Hamdi WS, Nasrallah GK, El Zowalaty ME, Al Khodor S, et al. Human microbiome and its association with health and diseases. J Cell Physiol. 2016; 231:1688–1694.10. Green BJ, Wiriyachaiporn S, Grainge C, Rogers GB, Kehagia V, Lau L, et al. Potentially pathogenic airway bacteria and neutrophilic inflammation in treatment resistant severe asthma. PLoS One. 2014; 9:e100645.11. Marri PR, Stern DA, Wright AL, Billheimer D, Martinez FD. Asthma-associated differences in microbial composition of induced sputum. J Allergy Clin Immunol. 2013; 131:346–352.12. Simpson JL, Daly J, Baines KJ, Yang IA, Upham JW, Reynolds PN, et al. Airway dysbiosis: Haemophilus influenzae and Tropheryma in poorly controlled asthma. Eur Respir J. 2016; 47:792–800.13. National Asthma. Expert Panel Report 3 (EPR-3): guidelines for the diagnosis and management of asthma: summary report 2007. J Allergy Clin Immunol. 2007; 120:5 Suppl. S94–138.14. Sohn SW, Lee HS, Park HW, Chang YS, Kim YK, Cho SH, et al. Evaluation of cytokine mRNA in induced sputum from patients with allergic rhinitis: relationship to airway hyperresponsiveness. Allergy. 2008; 63:268–273.15. Marsh TL. Terminal restriction fragment length polymorphism (T-RFLP): an emerging method for characterizing diversity among homologous populations of amplification products. Curr Opin Microbiol. 1999; 2:323–327.16. Schutte UM, Abdo Z, Bent SJ, Shyu C, Williams CJ, Pierson JD, et al. Advances in the use of terminal restriction fragment length polymorphism (T-RFLP) analysis of 16S rRNA genes to characterize microbial communities. Appl Microbiol Biotechnol. 2008; 80:365–380.17. Andoh A, Imaeda H, Aomatsu T, Inatomi O, Bamba S, Sasaki M, et al. Comparison of the fecal microbiota profiles between ulcerative colitis and Crohn’s disease using terminal restriction fragment length polymorphism analysis. J Gastroenterol. 2011; 46:479–486.18. Carroll IM, Ringel-Kulka T, Keku TO, Chang YH, Packey CD, Sartor RB, et al. Molecular analysis of the luminal- and mucosal-associated intestinal microbiota in diarrhea-predominant irritable bowel syndrome. Am J Physiol Gastrointest Liver Physiol. 2011; 301:G799–G807.19. Andoh A, Sakata S, Koizumi Y, Mitsuyama K, Fujiyama Y, Benno Y. Terminal restriction fragment length polymorphism analysis of the diversity of fecal microbiota in patients with ulcerative colitis. Inflamm Bowel Dis. 2007; 13:955–962.20. Andoh A, Tsujikawa T, Sasaki M, Mitsuyama K, Suzuki Y, Matsui T, et al. Faecal microbiota profile of Crohn’s disease determined by terminal restriction fragment length polymorphism analysis. Aliment Pharmacol Ther. 2009; 29:75–82.21. Matsumoto M, Sakamoto M, Hayashi H, Benno Y. Novel phylogenetic assignment database for terminal-restriction fragment length polymorphism analysis of human colonic microbiota. J Microbiol Methods. 2005; 61:305–319.22. Rogers GB, Carroll MP, Serisier DJ, Hockey PM, Jones G, Bruce KD. characterization of bacterial community diversity in cystic fibrosis lung infections by use of 16s ribosomal DNA terminal restriction fragment length polymorphism profiling. J Clin Microbiol. 2004; 42:5176–5183.23. Rogers GB, Hart CA, Mason JR, Hughes M, Walshaw MJ, Bruce KD. Bacterial diversity in cases of lung infection in cystic fibrosis patients: 16S ribosomal DNA (rDNA) length heterogeneity PCR and 16S rDNA terminal restriction fragment length polymorphism profiling. J Clin Microbiol. 2003; 41:3548–3558.24. Cardenas PA, Cooper PJ, Cox MJ, Chico M, Arias C, Moffatt MF, et al. Upper airways microbiota in antibiotic-naive wheezing and healthy infants from the tropics of rural Ecuador. PLoS One. 2012; 7:e46803.25. Park H, Shin JW, Park SG, Kim W. Microbial communities in the upper respiratory tract of patients with asthma and chronic obstructive pulmonary disease. PLoS One. 2014; 9:e109710.26. Venkataraman A, Bassis CM, Beck JM, Young VB, Curtis JL, Huffnagle GB, et al. Application of a neutral community model to assess structuring of the human lung microbiome. MBio. 2015; 6:e02284–e02214.27. Russell SL, Gold MJ, Hartmann M, Willing BP, Thorson L, Wlodarska M, et al. Early life antibiotic-driven changes in microbiota enhance susceptibility to allergic asthma. EMBO Rep. 2012; 13:440–447.28. Fortenberry JD. The uses of race and ethnicity in human microbiome research. Trends Microbiol. 2013; 21:165–166.29. Carroll IM, Chang YH, Park J, Sartor RB, Ringel Y. Luminal and mucosal-associated intestinal microbiota in patients with diarrhea-predominant irritable bowel syndrome. Gut Pathog. 2010; 2:19.30. Abrahamsson TR, Jakobsson HE, Andersson AF, Bjorksten B, Engstrand L, Jenmalm MC. Low gut microbiota diversity in early infancy precedes asthma at school age. Clin Exp Allergy. 2014; 44:842–850.31. Webley WC, Aldridge KL. Infectious asthma triggers: time to revise the hygiene hypothesis? Trends Microbiol. 2015; 23:389–391.32. Huang YJ, Nelson CE, Brodie EL, Desantis TZ, Baek MS, Liu J, et al. Airway microbiota and bronchial hyperresponsiveness in patients with suboptimally controlled asthma. J Allergy Clin Immunol. 2011; 127:372–381.33. Goleva E, Jackson LP, Harris JK, Robertson CE, Sutherland ER, Hall CF, et al. The effects of airway microbiome on corticosteroid responsiveness in asthma. Am J Respir Crit Care Med. 2013; 188:1193–1201.34. Hilty M, Burke C, Pedro H, Cardenas P, Bush A, Bossley C, et al. Disordered microbial communities in asthmatic airways. PLoS One. 2010; 5:e8578.35. Perez-Losada M, Castro-Nallar E, Bendall ML, Freishtat RJ, Crandall KA. Dual transcriptomic profiling of host and microbiota during health and disease in pediatric asthma. PLoS One. 2015; 10:e0131819.36. Chotirmall SH, Burke CM. Aging and the microbiome: implications for asthma in the elderly? Expert Rev Respir Med. 2015; 9:125–128.37. Trompette A, Gollwitzer ES, Yadava K, Sichelstiel AK, Sprenger N, Ngom-Bru C, et al. Gut microbiota metabolism of dietary fiber influences allergic airway disease and hematopoiesis. Nat Med. 2014; 20:159–166.38. Vital M, Harkema JR, Rizzo M, Tiedje J, Brandenberger C. Alterations of the murine gut microbiome with age and allergic airway disease. J Immunol Res. 2015; 2015:892568.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- The Usefulness of Induced Sputum Test in Asthma
- Interactions between NCR + ILC3s and the Microbiome in the Airways Shape Asthma Severity
- Inflammatory cells in the induced sputum of asthmatic patients
- Clinical characteristics and long-term outcomes related to sputum eosinophilia in Korean asthmatics
- Humoral immune responses in the induced sputum of asthmatic patients