J Breast Cancer.
2020 Feb;23(1):36-46. 10.4048/jbc.2020.23.e12.
Downregulation of N-myc and STAT Interactor Protein Predicts Aggressive Tumor Behavior and Poor Prognosis in Invasive Ductal Carcinoma
- Affiliations
-
- 1Department of Pathology, Design Hospital, Jeonju, Korea.
- 2Department of Pathology, Jeju National University Hospital, Jeju, Korea.
- 3Department of Pathology, Bucheon St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Bucheon, Korea.
- 4Department of Pathology, Seoul National University Hospital, Seoul National University College of Medicine, Seoul, Korea. nash77@snu.ac.kr
- KMID: 2470880
- DOI: http://doi.org/10.4048/jbc.2020.23.e12
Abstract
- PURPOSE
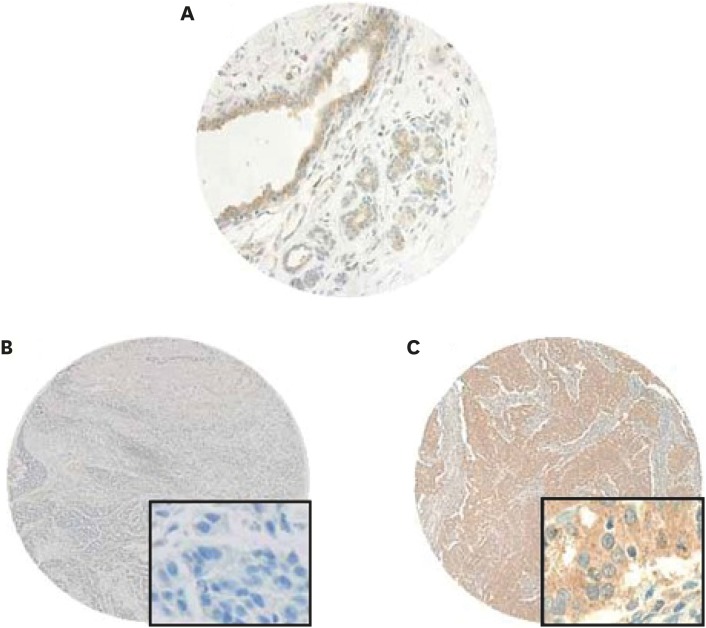
We investigated the expression of the N-myc and STAT interactor (NMI) protein in invasive ductal carcinoma tissue and estimated its clinicopathologic significance as a prognostic factor. The expression levels and prognostic significance of NMI were also analyzed according to the molecular subgroup of breast cancers.
METHODS
Human NMI detection by immunohistochemistry was performed using tissue microarrays of 382 invasive ductal carcinomas. The correlation of NMI expression with patient clinicopathological parameters and prognostic significance was analyzed and further assessed according to the molecular subgroup of breast cancers. Moreover, in vitro experiments with 13 breast cancer cell lines were carried out. We also validated NMI expression significance in The Cancer Genome Atlas cohort using the Human Protein Atlas (HPA) database.
RESULTS
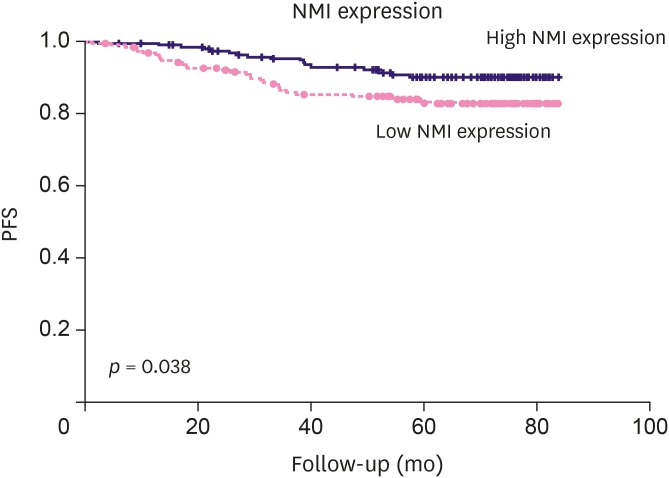
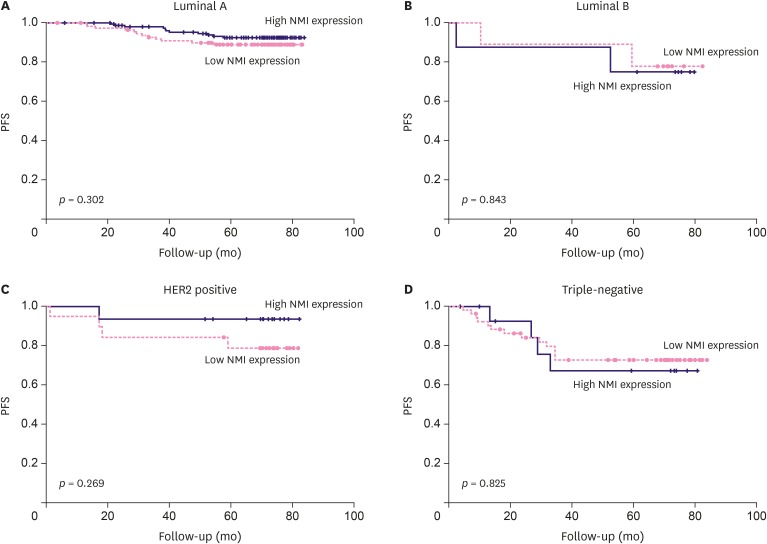
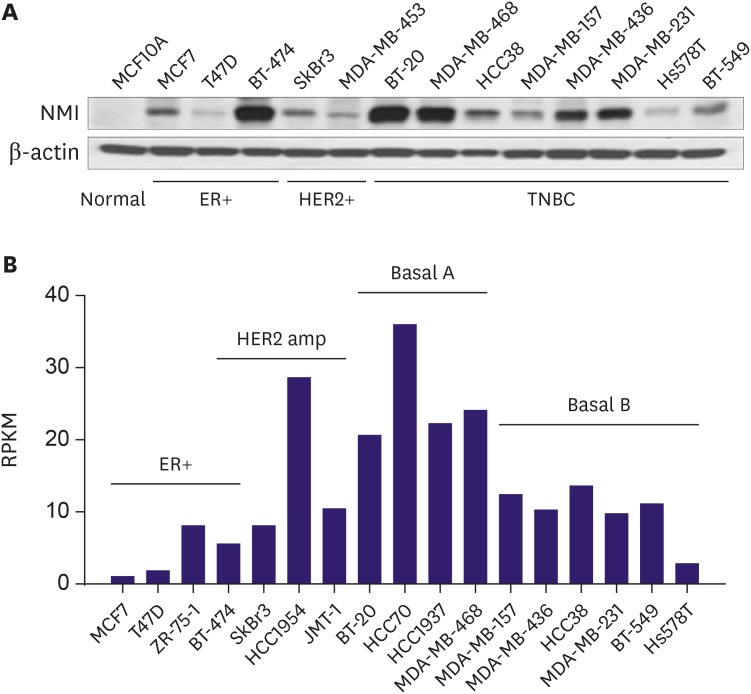
Low NMI expression was observed in 190 cases (49.7%). Low NMI expression was significantly associated with the "triple-negative" molecular subtype (p < 0.001), high nuclear grade (p < 0.001), high histologic grade (p < 0.001), and advanced anatomic stage (p = 0.041). Patients with low NMI expression had poorer progression-free survival (p = 0.038) than patients with high NMI expression. Low NMI expression was not significantly associated with patient prognosis in the molecular subgroup analysis. In vitro, a reduction of NMI expression was observed in 8 breast cancer cell lines, especially in the estrogen receptor-positive and basal B type of triple-negative breast cancer molecular subgroups. The HPA database showed that low NMI expression levels were associated with a lower survival probability compared with that associated with high NMI expression (p = 0.053).
CONCLUSION
NMI expression could be a useful prognostic biomarker and a potential novel therapeutic target in invasive ductal carcinoma.
MeSH Terms
Figure
Reference
-
1. Siegel RL, Miller KD, Jemal A. Cancer statistics, 2019. CA Cancer J Clin. 2019; 69:7–34. PMID: 30620402.
Article2. DeSantis CE, Ma J, Goding Sauer A, Newman LA, Jemal A. Breast cancer statistics, 2017, racial disparity in mortality by state. CA Cancer J Clin. 2017; 67:439–448. PMID: 28972651.
Article3. PDQ Adult Treatment Editorial Board. Breast cancer treatment (PDQ®): patient version. National Cancer Institute. PDQ Cancer Information Summaries. Bethesda (MD): National Cancer Institute;2002.4. Bao J, Zervos AS. Isolation and characterization of Nmi, a novel partner of Myc proteins. Oncogene. 1996; 12:2171–2176. PMID: 8668343.5. Chen J, Shpall RL, Meyerdierks A, Hagemeier M, Böttger EC, Naumovski L. Interferon-inducible Myc/STAT-interacting protein Nmi associates with IFP 35 into a high molecular mass complex and inhibits proteasome-mediated degradation of IFP 35. J Biol Chem. 2000; 275:36278–36284. PMID: 10950963.
Article6. Schlierf B, Lang S, Kosian T, Werner T, Wegner M. The high-mobility group transcription factor Sox10 interacts with the N-myc-interacting protein Nmi. J Mol Biol. 2005; 353:1033–1042. PMID: 16214168.
Article7. Zhang K, Zheng G, Yang YC. Stability of Nmi protein is controlled by its association with Tip60. Mol Cell Biochem. 2007; 303:1–8. PMID: 17406968.
Article8. Zhu M, John S, Berg M, Leonard WJ. Functional association of Nmi with Stat5 and Stat1 in IL-2- and IFNgamma-mediated signaling. Cell. 1999; 96:121–130. PMID: 9989503.9. Meng D, Chen Y, Yun D, Zhao Y, Wang J, Xu T, et al. High expression of N-myc (and STAT) interactor predicts poor prognosis and promotes tumor growth in human glioblastoma. Oncotarget. 2015; 6:4901–4919. PMID: 25669971.
Article10. Zhao J, Dong QZ, Zhong F, Cai LL, Qin ZY, Liu Y, et al. NMI promotes hepatocellular carcinoma progression via BDKRB2 and MAPK/ERK pathway. Oncotarget. 2017; 8:12174–12185. PMID: 28077802.
Article11. Li Z, Hou J, Sun L, Wen T, Wang L, Zhao X, et al. NMI mediates transcription-independent ARF regulation in response to cellular stresses. Mol Biol Cell. 2012; 23:4635–4646. PMID: 23034180.
Article12. Fillmore RA, Mitra A, Xi Y, Ju J, Scammell J, Shevde LA, et al. Nmi (N-Myc interactor) inhibits Wnt/beta-catenin signaling and retards tumor growth. Int J Cancer. 2009; 125:556–564. PMID: 19358268.13. Devine DJ, Rostas JW, Metge BJ, Das S, Mulekar MS, Tucker JA, et al. Loss of N-Myc interactor promotes epithelial-mesenchymal transition by activation of TGF-β/SMAD signaling. Oncogene. 2014; 33:2620–2628. PMID: 23770854.
Article14. Pruitt HC, Metge BJ, Weeks SE, Chen D, Wei S, Kesterson RA, et al. Conditional knockout of N-Myc and STAT interactor disrupts normal mammary development and enhances metastatic ability of mammary tumors. Oncogene. 2018; 37:1610–1623. PMID: 29326438.
Article15. Rostas JW 3rd, Pruitt HC, Metge BJ, Mitra A, Bailey SK, Bae S, et al. microRNA-29 negatively regulates EMT regulator N-myc interactor in breast cancer. Mol Cancer. 2014; 13:200. PMID: 25174825.
Article16. Metge BJ, Mitra A, Chen D, Shevde LA, Samant RS. N-Myc and STAT interactor regulates autophagy and chemosensitivity in breast cancer cells. Sci Rep. 2015; 5:11995. PMID: 26146406.
Article17. Lakhani SR, Ellis IO, Schnitt SJ, Tan PH, van de Vijver MJ. WHO Classification of Tumours of the Breast. Lyon: International Agency for Research on Cancer;2012.18. Edge SB, Greene FL, Byrd DR, Brookland RK, Washington MK, Gershenwald JE, et al. AJCC Cancer Staging Manual. New York (NY): Springer;2017.19. Elston CW, Ellis IO. Pathological prognostic factors in breast cancer. I. The value of histological grade in breast cancer: experience from a large study with long-term follow-up. Histopathology. 2002; 41:154–161. PMID: 12405947.
Article20. Goldhirsch A, Wood WC, Coates AS, Gelber RD, Thürlimann B, Senn HJ, et al. Strategies for subtypes--dealing with the diversity of breast cancer: highlights of the St. Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2011. Ann Oncol. 2011; 22:1736–1747. PMID: 21709140.
Article21. Hammond ME, Hayes DF, Dowsett M, Allred DC, Hagerty KL, Badve S, et al. American Society of Clinical Oncology/College of American Pathologists guideline recommendations for immunohistochemical testing of estrogen and progesterone receptors in breast cancer. J Clin Oncol. 2010; 28:2784–2795. PMID: 20404251.
Article22. Wolff AC, Hammond ME, Hicks DG, Dowsett M, McShane LM, Allison KH, et al. Recommendations for human epidermal growth factor receptor 2 testing in breast cancer: American Society of Clinical Oncology/College of American Pathologists clinical practice guideline update. J Clin Oncol. 2013; 31:3997–4013. PMID: 24101045.
Article23. Dai X, Cheng H, Bai Z, Li J. Breast cancer cell line classification and its relevance with breast tumor subtyping. J Cancer. 2017; 8:3131–3141. PMID: 29158785.
Article24. Li H, Lee TH, Avraham H. A novel tricomplex of BRCA1, Nmi, and c-Myc inhibits c-Myc-induced human telomerase reverse transcriptase gene (hTERT) promoter activity in breast cancer. J Biol Chem. 2002; 277:20965–20973. PMID: 11916966.
Article25. Hollier BG, Evans K, Mani SA. The epithelial-to-mesenchymal transition and cancer stem cells: a coalition against cancer therapies. J Mammary Gland Biol Neoplasia. 2009; 14:29–43. PMID: 19242781.
Article26. Thiery JP, Sleeman JP. Complex networks orchestrate epithelial-mesenchymal transitions. Nat Rev Mol Cell Biol. 2006; 7:131–142. PMID: 16493418.
Article27. Tang X, Ding CK, Wu J, Sjol J, Wardell S, Spasojevic I, et al. Cystine addiction of triple-negative breast cancer associated with EMT augmented death signaling. Oncogene. 2017; 36:4235–4242. PMID: 27869167.
Article28. Lohia M, Qin Y, Macara IG. The Scribble polarity protein stabilizes E-cadherin/p120-catenin binding and blocks retrieval of E-cadherin to the Golgi. PLoS One. 2012; 7:e51130. PMID: 23226478.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Fine Needle Aspiration Cytology of Invasive Micropapillary Carcinoma of the Breast
- The Glycolytic Phenotype is Correlated with Aggressiveness and Poor Prognosis in Invasive Ductal Carcinomas
- A Case of Bilat. Choroidal Metastasis of Breast Invasive Ductal Carcinoma
- The Difference of Cathepsin D Expression between Invasive Ductal Carcinoma and Ductal Carcinoma In Situ of the Breast
- Expression of p53 and Rb Proteins in Invasive Ductal Carcinoma of the Breast