Diabetes Metab J.
2019 Oct;43(5):649-658. 10.4093/dmj.2018.0232.
Essential Role of Protein Arginine Methyltransferase 1 in Pancreas Development by Regulating Protein Stability of Neurogenin 3
- Affiliations
-
- 1Graduate School of Medical Science and Engineering, Korea Advanced Institute of Science and Technology, Daejeon, Korea. hailkim@kaist.edu
- 2Department of Internal Medicine, CHA Bundang Medical Center, CHA University, Seongnam, Korea.
- 3Division of Life Sciences, Korea University, Seoul, Korea.
- 4Department of Pathology, Daejeon St. Mary's Hospital, College of Medicine, The Catholic University of Korea, Daejeon, Korea.
- 5KAIST Institute for the BioCentury, Korea Advanced Institute of Science and Technology, Daejeon, Korea.
- KMID: 2460958
- DOI: http://doi.org/10.4093/dmj.2018.0232
Abstract
- BACKGROUND
Protein arginine methyltransferase 1 (PRMT1) is a major enzyme responsible for the formation of methylarginine in mammalian cells. Recent studies have revealed that PRMT1 plays important roles in the development of various tissues. However, its role in pancreas development has not yet been elucidated.
METHODS
Pancreatic progenitor cell-specific Prmt1 knock-out (Prmt1 PKO) mice were generated and characterized for their metabolic and histological phenotypes and their levels of Neurog3 gene expression and neurogenin 3 (NGN3) protein expression. Protein degradation assays were performed in mPAC cells.
RESULTS
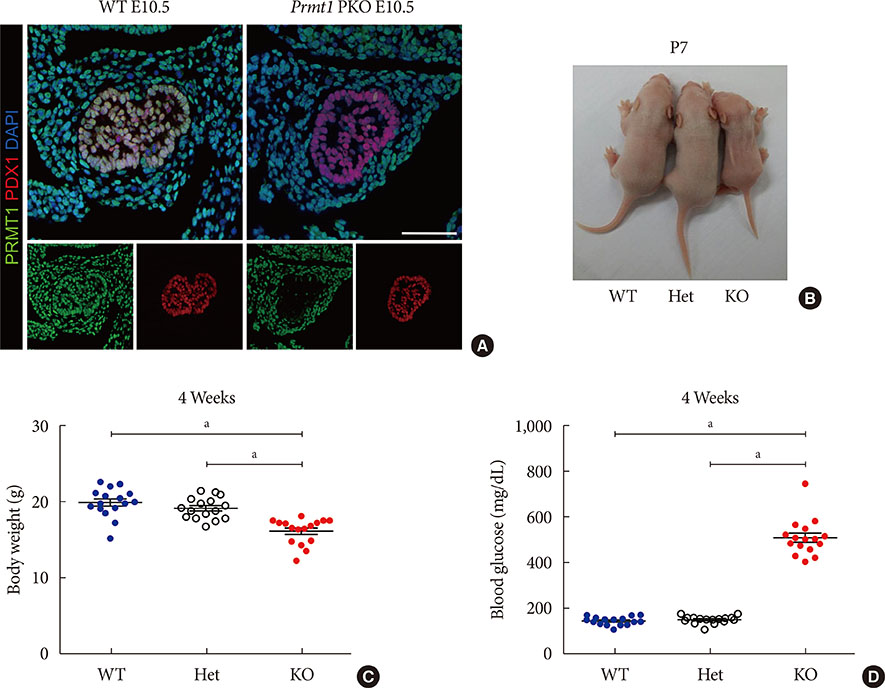
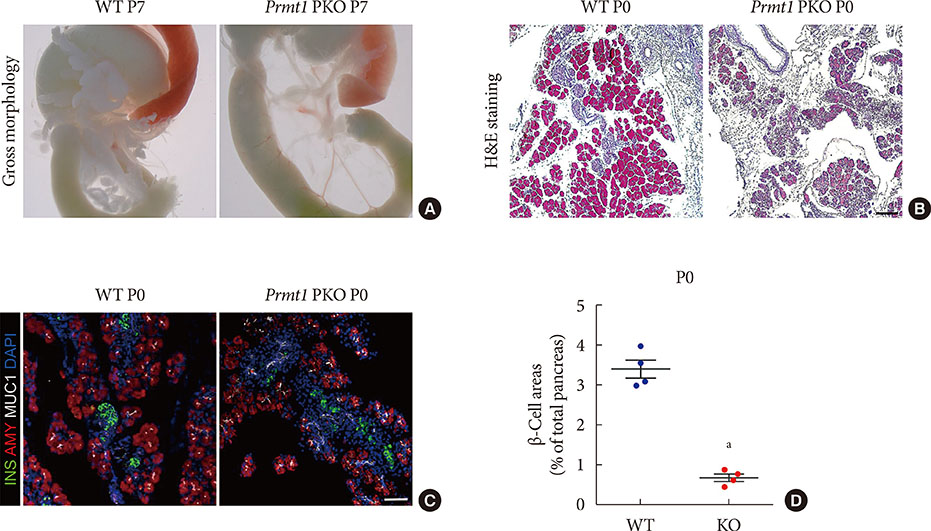
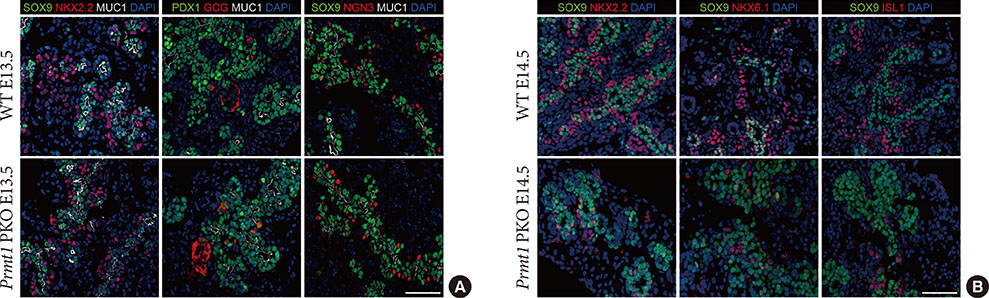
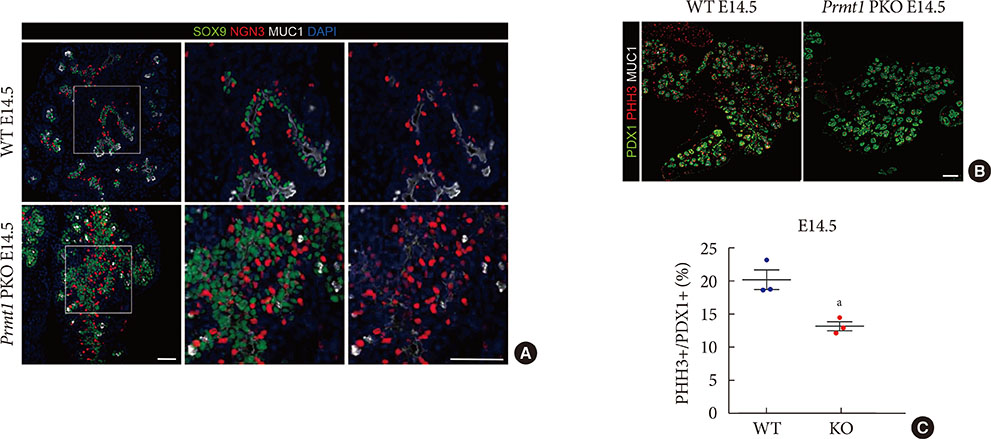
Prmt1 PKO mice showed growth retardation and a severely diabetic phenotype. The pancreatic size and β-cell mass were significantly reduced in Prmt1 PKO mice. Proliferation of progenitor cells during the secondary transition was decreased and endocrine cell differentiation was impaired. These defects in pancreas development could be attributed to the sustained expression of NGN3 in progenitor cells. Protein degradation assays in mPAC cells revealed that PRMT1 was required for the rapid degradation of NGN3.
CONCLUSION
PRMT1 critically contributes to pancreas development by destabilizing the NGN3 protein.
Keyword
MeSH Terms
Figure
Reference
-
1. Bedford MT, Clarke SG. Protein arginine methylation in mammals: who, what, and why. Mol Cell. 2009; 33:1–13.2. Wei H, Mundade R, Lange KC, Lu T. Protein arginine methylation of non-histone proteins and its role in diseases. Cell Cycle. 2014; 13:32–41.3. Kleinschmidt MA, Streubel G, Samans B, Krause M, Bauer UM. The protein arginine methyltransferases CARM1 and PRMT1 cooperate in gene regulation. Nucleic Acids Res. 2008; 36:3202–3213.4. Ohkura N, Takahashi M, Yaguchi H, Nagamura Y, Tsukada T. Coactivator-associated arginine methyltransferase 1, CARM1, affects pre-mRNA splicing in an isoform-specific manner. J Biol Chem. 2005; 280:28927–28935.5. Cha B, Kim W, Kim YK, Hwang BN, Park SY, Yoon JW, Park WS, Cho JW, Bedford MT, Jho EH. Methylation by protein arginine methyltransferase 1 increases stability of Axin, a negative regulator of Wnt signaling. Oncogene. 2011; 30:2379–2389.6. Blanc RS, Richard S. Arginine methylation: the coming of age. Mol Cell. 2017; 65:8–24.7. Tang J, Frankel A, Cook RJ, Kim S, Paik WK, Williams KR, Clarke S, Herschman HR. PRMT1 is the predominant type I protein arginine methyltransferase in mammalian cells. J Biol Chem. 2000; 275:7723–7730.8. Wada K, Inoue K, Hagiwara M. Identification of methylated proteins by protein arginine N-methyltransferase 1, PRMT1, with a new expression cloning strategy. Biochim Biophys Acta. 2002; 1591:1–10.9. Pawlak MR, Scherer CA, Chen J, Roshon MJ, Ruley HE. Arginine N-methyltransferase 1 is required for early postimplantation mouse development, but cells deficient in the enzyme are viable. Mol Cell Biol. 2000; 20:4859–4869.10. Hashimoto M, Murata K, Ishida J, Kanou A, Kasuya Y, Fukamizu A. Severe hypomyelination and developmental defects are caused in mice lacking protein arginine methyltransferase 1 (PRMT1) in the central nervous system. J Biol Chem. 2016; 291:2237–2245.11. Blanc RS, Vogel G, Li X, Yu Z, Li S, Richard S. Arginine methylation by PRMT1 regulates muscle stem cell fate. Mol Cell Biol. 2017; 37:e00457-16.12. Roder PV, Wu B, Liu Y, Han W. Pancreatic regulation of glucose homeostasis. Exp Mol Med. 2016; 48:e219.13. Burlison JS, Long Q, Fujitani Y, Wright CV, Magnuson MA. Pdx-1 and Ptf1a concurrently determine fate specification of pancreatic multipotent progenitor cells. Dev Biol. 2008; 316:74–86.14. Oliver-Krasinski JM, Stoffers DA. On the origin of the beta cell. Genes Dev. 2008; 22:1998–2021.15. Choi D, Oh KJ, Han HS, Yoon YS, Jung CY, Kim ST, Koo SH. Protein arginine methyltransferase 1 regulates hepatic glucose production in a FoxO1-dependent manner. Hepatology. 2012; 56:1546–1556.16. Gu G, Dubauskaite J, Melton DA. Direct evidence for the pancreatic lineage: NGN3+ cells are islet progenitors and are distinct from duct progenitors. Development. 2002; 129:2447–2457.17. Fischer AH, Jacobson KA, Rose J, Zeller R. Hematoxylin and eosin staining of tissue and cell sections. CSH Protoc. 2008; 2008:pdb.prot4986.18. Livak KJ, Schmittgen TD. Analysis of relative gene expression data using real-time quantitative PCR and the 2(-Delta Delta C(T)) method. Methods. 2001; 25:402–408.19. Pan FC, Wright C. Pancreas organogenesis: from bud to plexus to gland. Dev Dyn. 2011; 240:530–565.20. Gradwohl G, Dierich A, LeMeur M, Guillemot F. Neurogenin3 is required for the development of the four endocrine cell lineages of the pancreas. Proc Natl Acad Sci U S A. 2000; 97:1607–1611.21. Lee JC, Smith SB, Watada H, Lin J, Scheel D, Wang J, Mirmira RG, German MS. Regulation of the pancreatic pro-endocrine gene neurogenin3. Diabetes. 2001; 50:928–936.22. Johansson KA, Dursun U, Jordan N, Gu G, Beermann F, Gradwohl G, Grapin-Botton A. Temporal control of neurogenin3 activity in pancreas progenitors reveals competence windows for the generation of different endocrine cell types. Dev Cell. 2007; 12:457–465.23. Apelqvist A, Li H, Sommer L, Beatus P, Anderson DJ, Honjo T, Hrabe de Angelis M, Lendahl U, Edlund H. Notch signalling controls pancreatic cell differentiation. Nature. 1999; 400:877–881.24. Bankaitis ED, Bechard ME, Wright CV. Feedback control of growth, differentiation, and morphogenesis of pancreatic endocrine progenitors in an epithelial plexus niche. Genes Dev. 2015; 29:2203–2216.25. Rukstalis JM, Habener JF. Neurogenin3: a master regulator of pancreatic islet differentiation and regeneration. Islets. 2009; 1:177–184.26. Bechard ME, Bankaitis ED, Hipkens SB, Ustione A, Piston DW, Yang YP, Magnuson MA, Wright CV. Precommitment low-level Neurog3 expression defines a long-lived mitotic endocrine-biased progenitor pool that drives production of endocrine-committed cells. Genes Dev. 2016; 30:1852–1865.27. Yu XX, Qiu WL, Yang L, Li LC, Zhang YW, Xu CR. Dynamics of chromatin marks and the role of JMJD3 during pancreatic endocrine cell fate commitment. Development. 2018; 145:dev163162.28. Krentz NAJ, van Hoof D, Li Z, Watanabe A, Tang M, Nian C, German MS, Lynn FC. Phosphorylation of NEUROG3 links endocrine differentiation to the cell cycle in pancreatic progenitors. Dev Cell. 2017; 41:129–142.29. Azzarelli R, Hurley C, Sznurkowska MK, Rulands S, Hardwick L, Gamper I, Ali F, McCracken L, Hindley C, McDuff F, Nestorowa S, Kemp R, Jones K, Gottgens B, Huch M, Evan G, Simons BD, Winton D, Philpott A. Multi-site neurogenin3 phosphorylation controls pancreatic endocrine differentiation. Dev Cell. 2017; 41:274–286.30. Huang S, Litt M, Felsenfeld G. Methylation of histone H4 by arginine methyltransferase PRMT1 is essential in vivo for many subsequent histone modifications. Genes Dev. 2005; 19:1885–1893.31. Xu J, Wang AH, Oses-Prieto J, Makhijani K, Katsuno Y, Pei M, Yan L, Zheng YG, Burlingame A, Bruckner K, Derynck R. Arginine methylation initiates BMP-induced smad signaling. Mol Cell. 2013; 51:5–19.32. Zhang L, Tran NT, Su H, Wang R, Lu Y, Tang H, Aoyagi S, Guo A, Khodadadi-Jamayran A, Zhou D, Qian K, Hricik T, Cote J, Han X, Zhou W, Laha S, Abdel-Wahab O, Levine RL, Raffel G, Liu Y, Chen D, Li H, Townes T, Wang H, Deng H, Zheng YG, Leslie C, Luo M, Zhao X. Cross-talk between PRMT1-mediated methylation and ubiquitylation on RBM15 controls RNA splicing. Elife. 2015; 4:e07938.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Identification of Protein Arginine Methyltransferase 5 as a Regulator for Encystation of Acanthamoeba
- Identification and Characterization of Protein Arginine Methyltransferase 1 in Acanthamoeba castellanii
- Arginine methylation as a key post-translational modification in skeletal muscle homeostasis: a review
- Roles of Protein Arginine Methyltransferases in the Control of Glucose Metabolism
- Increased methylation of the cytosolic 20-kD protein is accompanied by liver regeneration in a hepatectomized rat