Cancer Res Treat.
2022 Oct;54(4):970-984. 10.4143/crt.2021.1078.
Reclassification of Five BRCA1/2 Variants with Unknown Significance Using Complex Functional Study
- Affiliations
-
- 1Department of Molecular Genetics, National Institute of Oncology, Budapest, Hungary
- 2Hereditary Cancers Research Group, Hungarian Academy of Sciences - Semmelweis University, Budapest, Hungary
- KMID: 2534177
- DOI: http://doi.org/10.4143/crt.2021.1078
Abstract
- Purpose
While BRCA1/2 genes are commonly investigated, variants of unknown significance (VUS) and variants with potential splice effect are still being detected and they represent a substantial challenge in genetic counseling and therapy.
Materials and Methods
Out of genetically tested 3,568 hereditary breast and ovarian cancer probands five, functionally not investigated variants with potential splice-modifying effect were subjected to functional characterization. Transcript-level analysis on peripheral blood-derived RNA of the carriers was performed to test aberrant splicing. The completeness of the aberrant splicing event was also studied, existence and extent of nonsense-mediated decay was even addressed. Clinical and phenotype data, pedigree and co-segregation analyses were also done. Locus-specific loss of heterozygosity (LOH) in tumor tissues was additionally tested.
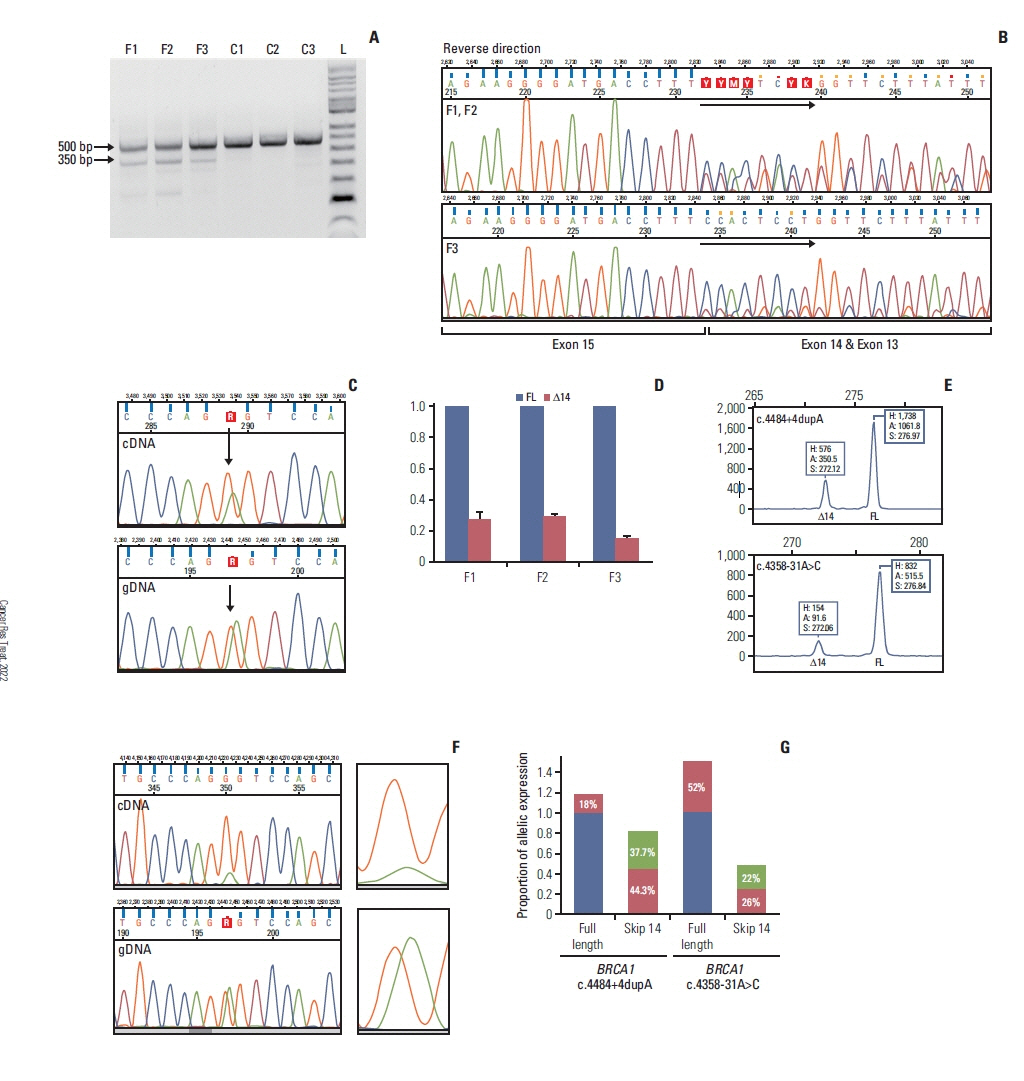
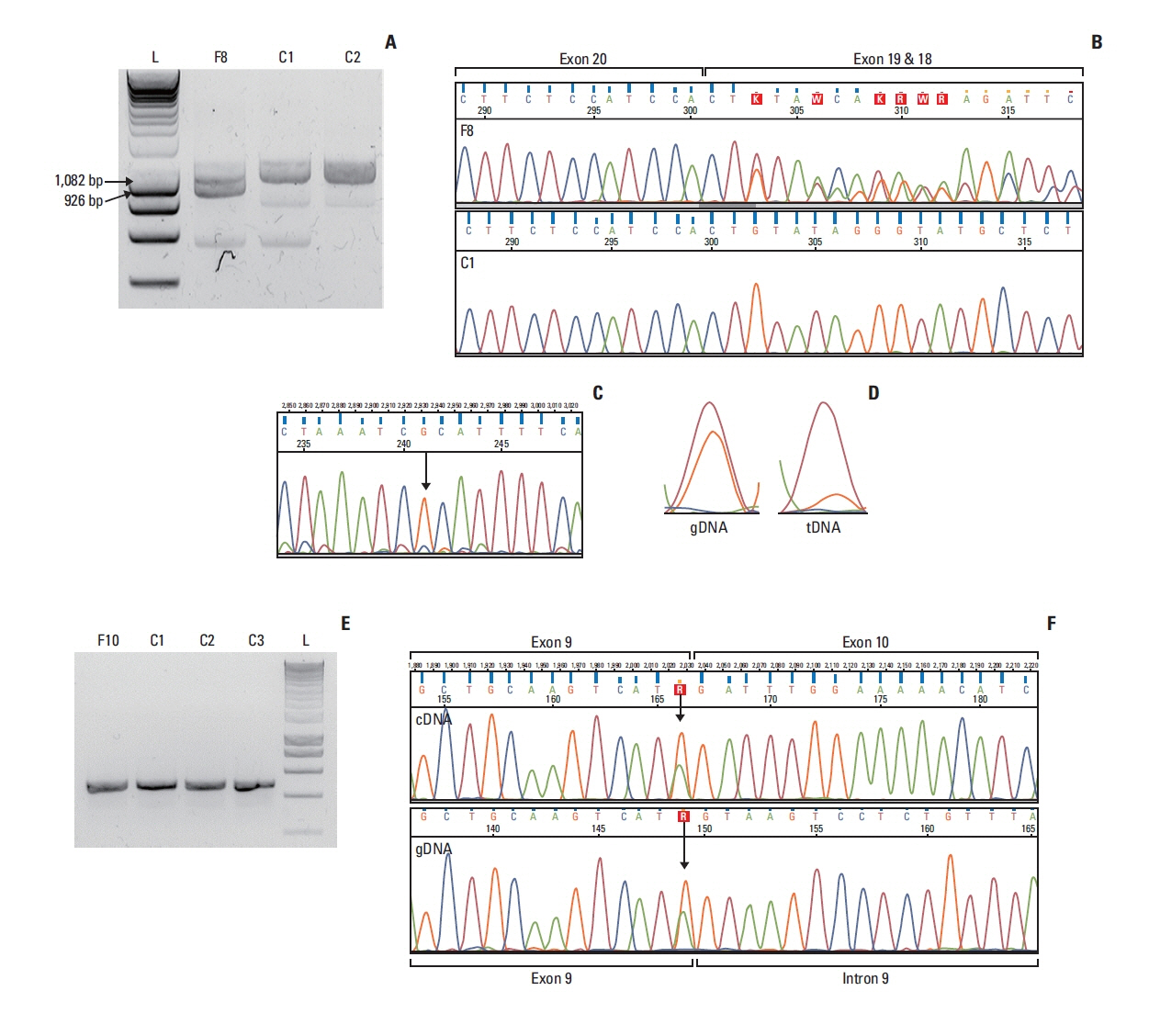
Results
In case of the BRCA1:c.4484+4dupA and the BRCA1:c.5407-10G>A variants functional results allowed us to reclassify them from VUS into likely pathogenic category. BRCA1:c.4358-31A>C, by producing incomplete aberrant splicing, was highlighted as strong VUS, but in lack of other supporting evidence, re-categorization was not possible. The likely pathogenic assertion of previously not reported BRCA2:c.8487G>T was reinforced based on its spliceogenic property and tumor LOH, while BRCA2:c.793G>A failed to present aberrant splicing in spite of suggestive predictions, which altered its original VUS evaluation into likely benign class.
Conclusion
We presented molecular and clinical evidence for reclassification of four out of five BRCA1/2 variants. Both up- and down-classification harbour important clinical significance. Patients carrying re-classified pathogenic variants in the future will not be dropped out from medical surveillance, preventive measures, treatment and predictive family screening in relatives at risk.
Figure
Reference
-
References
1. Robson M, Offit K. Clinical practice. Management of an inherited predisposition to breast cancer. N Engl J Med. 2007; 357:154–62.
Article2. Federici G, Soddu S. Variants of uncertain significance in the era of high-throughput genome sequencing: a lesson from breast and ovary cancers. J Exp Clin Cancer Res. 2020; 39:46.
Article3. Singer CF, Balmana J, Burki N, Delaloge S, Filieri ME, Gerdes AM, et al. Genetic counselling and testing of susceptibility genes for therapeutic decision-making in breast cancer-an European consensus statement and expert recommendations. Eur J Cancer. 2019; 106:54–60.
Article4. Richards S, Aziz N, Bale S, Bick D, Das S, Gastier-Foster J, et al. Standards and guidelines for the interpretation of sequence variants: a joint consensus recommendation of the American College of Medical Genetics and Genomics and the Association for Molecular Pathology. Genet Med. 2015; 17:405–24.
Article5. Kuchenbaecker KB, Hopper JL, Barnes DR, Phillips KA, Mooij TM, Roos-Blom MJ, et al. Risks of breast, ovarian, and contralateral breast cancer for BRCA1 and BRCA2 mutation carriers. JAMA. 2017; 317:2402–16.6. Parsons MT, Tudini E, Li H, Hahnen E, Wappenschmidt B, Feliubadalo L, et al. Large scale multifactorial likelihood quantitative analysis of BRCA1 and BRCA2 variants: an ENIGMA resource to support clinical variant classification. Hum Mutat. 2019; 40:1557–78.7. Harper K. BRCA exchange launches. Cancer Discov. 2019; 9:311–2.
Article8. Murray ML, Cerrato F, Bennett RL, Jarvik GP. Follow-up of carriers of BRCA1 and BRCA2 variants of unknown significance: variant reclassification and surgical decisions. Genet Med. 2011; 13:998–1005.
Article9. Bevers TB, Anderson BO, Bonaccio E, Buys S, Daly MB, Dempsey PJ, et al. NCCN clinical practice guidelines in oncology: breast cancer screening and diagnosis. J Natl Compr Canc Netw. 2009; 7:1060–96.10. Deignan JL, Chung WK, Kearney HM, Monaghan KG, Rehder CW, Chao EC, et al. Points to consider in the reevaluation and reanalysis of genomic test results: a statement of the American College of Medical Genetics and Genomics (ACMG). Genet Med. 2019; 21:1267–70.
Article11. Orban TI, Olah E. Emerging roles of BRCA1 alternative splicing. Mol Pathol. 2003; 56:191–7.
Article12. Stoilov P, Meshorer E, Gencheva M, Glick D, Soreq H, Stamm S. Defects in pre-mRNA processing as causes of and predisposition to diseases. DNA Cell Biol. 2002; 21:803–18.
Article13. Marmolejo DH, Wong MYZ, Bajalica-Lagercrantz S, Tischkowitz M, Balmana J. Extended ERN-GENTURIS Thematic Group. Overview of hereditary breast and ovarian cancer (HBOC) guidelines across Europe. Eur J Med Genet. 2021; 64:104350.
Article14. Culp M, Johnson K, Michailides G. ada: an R package for stochastic boosting. J Stat Softw. 2006; 17:1–27.15. Breiman L. Random forests. Mach Learn. 2001; 45:5–32.16. Liu X, Li C, Mou C, Dong Y, Tu Y. dbNSFP v4: a comprehensive database of transcript-specific functional predictions and annotations for human nonsynonymous and splice-site SNVs. Genome Med. 2020; 12:103.
Article17. Nazari I, Tayara H, Chong KT. Branch point selection in RNA splicing using deep learning. IEEE Access. 2019; 7:1800–7.
Article18. Paggi JM, Bejerano G. A sequence-based, deep learning model accurately predicts RNA splicing branchpoints. RNA. 2018; 24:1647–58.
Article19. Papp J, Kovacs ME, Solyom S, Kasler M, Borresen-Dale AL, Olah E. High prevalence of germline STK11 mutations in Hungarian Peutz-Jeghers Syndrome patients. BMC Med Genet. 2010; 11:169.20. Kopanos C, Tsiolkas V, Kouris A, Chapple CE, Albarca Aguilera M, Meyer R, et al. VarSome: the human genomic variant search engine. Bioinformatics. 2019; 35:1978–80.
Article21. Goldgar DE, Easton DF, Deffenbaugh AM, Monteiro AN, Tavtigian SV, Couch FJ, et al. Integrated evaluation of DNA sequence variants of unknown clinical significance: application to BRCA1 and BRCA2. Am J Hum Genet. 2004; 75:535–44.
Article22. Wimmer K, Schamschula E, Wernstedt A, Traunfellner P, Amberger A, Zschocke J, et al. AG-exclusion zone revisited: lessons to learn from 91 intronic NF1 3′ splice site mutations outside the canonical AG-dinucleotides. Hum Mutat. 2020; 41:1145–56.
Article23. Findlay GM, Daza RM, Martin B, Zhang MD, Leith AP, Gasperini M, et al. Accurate classification of BRCA1 variants with saturation genome editing. Nature. 2018; 562:217–22.
Article24. Houdayer C, Caux-Moncoutier V, Krieger S, Barrois M, Bonnet F, Bourdon V, et al. Guidelines for splicing analysis in molecular diagnosis derived from a set of 327 combined in silico/in vitro studies on BRCA1 and BRCA2 variants. Hum Mutat. 2012; 33:1228–38.
Article25. Chehade R, Pettapiece-Phillips R, Salmena L, Kotlyar M, Jurisica I, Narod SA, et al. Reduced BRCA1 transcript levels in freshly isolated blood leukocytes from BRCA1 mutation carriers is mutation specific. Breast Cancer Res. 2016; 18:87.
Article26. de la Hoya M, Soukarieh O, Lopez-Perolio I, Vega A, Walker LC, van Ierland Y, et al. Combined genetic and splicing analysis of BRCA1 c.[594-2A>C; 641A>G] highlights the relevance of naturally occurring in-frame transcripts for developing disease gene variant classification algorithms. Hum Mol Genet. 2016; 25:2256–68.
Article27. Zhang L, Bacares R, Boyar S, Hudis C, Nafa K, Offit K. cDNA analysis demonstrates that the BRCA2 intronic variant IVS4-12del5 is a deleterious mutation. Mutat Res. 2009; 663:84–9.
Article28. Walsh MF, Ritter DI, Kesserwan C, Sonkin D, Chakravarty D, Chao E, et al. Integrating somatic variant data and biomarkers for germline variant classification in cancer predisposition genes. Hum Mutat. 2018; 39:1542–52.
Article29. Maxwell KN, Wubbenhorst B, Wenz BM, De Sloover D, Pluta J, Emery L, et al. BRCA locus-specific loss of heterozygosity in germline BRCA1 and BRCA2 carriers. Nat Commun. 2017; 8:319.
Article30. Bonnet C, Krieger S, Vezain M, Rousselin A, Tournier I, Martins A, et al. Screening BRCA1 and BRCA2 unclassified variants for splicing mutations using reverse transcription PCR on patient RNA and an ex vivo assay based on a splicing reporter minigene. J Med Genet. 2008; 45:438–46.
Article31. Mesman RL, Calleja F, de la Hoya M, Devilee P, van Asperen CJ, Vrieling H, et al. Alternative mRNA splicing can attenuate the pathogenicity of presumed loss-of-function variants in BRCA2. Genet Med. 2020; 22:1355–65.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Reclassification of BRCA1 and BRCA2 variants found in ovarian epithelial, fallopian tube, and primary peritoneal cancers
- Identification of a Novel BRCA1 Pathogenic Mutation in Korean Patients Following Reclassification of BRCA1 and BRCA2 Variants According to the ACMG Standards and Guidelines Using Relevant Ethnic Controls
- Applying Functional Assay Evidence to Interpret Sequence Variants Identified in Hereditary Cancer Genes
- Clinical significance of variants of unknown significances in BRCA genes
- Prognostic implications of ductal carcinoma in situ components in BRCA1/2-positive breast cancer: a retrospective cohort study