Nat Prod Sci.
2019 Sep;25(3):222-227. 10.20307/nps.2019.25.3.222.
Assessment of the Purity of Emodin by Quantitative Nuclear Magnetic Resonance Spectroscopy and Mass Balance
- Affiliations
-
- 1Department of Biosystems and Biotechnology, College of Life Sciences and Biotechnology, Korea University, Seoul 02841, Republic of Korea. dongholee@korea.ac.kr
- 2Graduate School of Pharmaceutical Sciences, College of Pharmacy, Ewha Womans University, Seoul 03760, Republic of Korea.
- 3Department of Environmental Science and Ecological Engineering, College of Life Sciences and Biotechnology, Korea University, Seoul 02841, Republic of Korea.
- KMID: 2459963
- DOI: http://doi.org/10.20307/nps.2019.25.3.222
Abstract
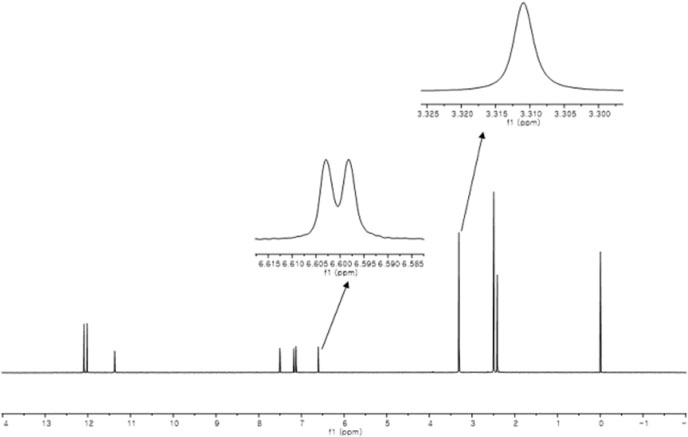
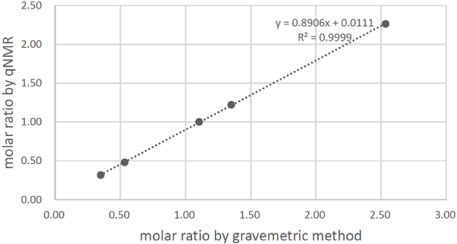
- Quantitative nuclear magnetic resonance (qNMR) is a well-established method adopted by international pharmacopoeia for quantitative and purity analyses. Emodin is a type of anthraquinone, well known as the main active component of Fabaceae, Polygonaceae and Rhamnaceae. Purity analysis of emodin is usually performed by using the high-performance liquid chromatography (HPLC)-UV method. However, it cannot detect impurities such as salts, volatile matter, and trace elements. Using the qNMR method, it is possible to determine the compound content as well as the nature of the impurities. Several experimental parameters were optimized for the quantification, such as relaxation delay, spectral width, number of scans, temperature, pulse width, and acquisition time. The method was validated, and the results of the qNMR method were compared with those obtained by the HPLC and mass balance analysis methods. The qNMR method is specific, rapid, simple, and therefore, a valuable and reliable method for the purity analysis of emodin.
Keyword
MeSH Terms
Figure
Reference
-
1. Malz F, Jancke H. J Pharm Biomed Anal. 2005; 38:813–823.2. Al Deen TS, Hibbert DB, Hook JM, Wells RJ. Anal Chim Acta. 2002; 474:125–135.3. Al Deen TS, Hibbert DB, Hook JM, Wells RJ. Accred Qual Assur. 2004; 9:55–63.4. Moura S, Carvalho FG, de Oliveira CDR, Pinto E, Yonamine M. Phytochem Lett. 2010; 3:79–83.5. Ohtsuki T, Sato K, Sugimoto N, Akiyama H, Kawamura Y. Talanta. 2012; 99:342–348.6. Izhaki I. New Phytol. 2002; 155:205–217.7. Cui YT, Liu B, Xie J, Xu P, Habte-Tsion HM, Zhang YY. Fish Shellfish Immunol. 2014; 38:74–79.8. Gupta SC, Prasad S, Aggarwal BB. Anti-inflammatory Nutraceuticals and Chronic Diseases. Springer: Basel;2016. p. 47–64.9. Li CL, Ma J, Zheng L, Li HJ, Li P. J Pharm Biomed Anal. 2012; 71:71–78.10. Simmler C, Napolitano JG, McAlpine JB, Chen SN, Pauli GF. Curr Opin Biotechnol. 2014; 25:51–59.11. Pauli GF. Phytochem Anal. 2001; 12:28–42.
Article12. Bharti SK, Roy R. TrAC Trends Anal Chem. 2012; 35:5–26.13. Saito T, Ihara T, Koike M, Kinugasa S, Fujimine Y, Nose K, Hirai T. Accred Qual Assur. 2009; 14:79–86.14. Wells R, Cheung J, Hook J. The use of qNMR for the analysis of agrochemicals. Elsevier: Amsterdam;2008. p. 291–315.
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Magnetic Resonance Spectroscopy
- Use of Nuclear Magnetic Resonance Spectroscopy in Analysis of Fennel Essential Oil
- Current Methods of Human Body Composition Assessment
- Comparison of in Vivo, in Vitro 3T MR Spectroscopy and Proton NMR Spectroscopy for the Fluid from Cystic Tumor: Preliminary Study
- Analysis of gallbladder bile by nuclear magnetic resonance (NMR) spectroscopy