Cancer Res Treat.
2023 Apr;55(2):659-670. 10.4143/crt.2022.1440.
Plasma Cell-Free DNA in Uterine Cervical Cancer: Therapeutic Monitoring and Prognostic Values after Radical Radiotherapy
- Affiliations
-
- 1Department of Radiation Oncology, Seoul National University College of Medicine, Seoul, Korea
- 2Department of Radiation Oncology, Soonchunhyang University Seoul Hospital, Seoul, Korea
- 3Transdisciplinary Department of Medicine and Advanced Technology, Seoul National University Hospital, Seoul, Korea
- 4Interdisciplinary Program in Bioengineering, Seoul National University, Seoul, Korea
- 5Department of Radiation Oncology, Kyung Hee University Hospital at Gangdong, Seoul, Korea
- 6Department of Radiation Oncology, Seoul National University Hospital, Seoul, Korea
- KMID: 2541253
- DOI: http://doi.org/10.4143/crt.2022.1440
Abstract
- Purpose
In the present study, we aimed to establish a liquid biopsy-based monitoring method using peripheral blood cell-free DNA (cfDNA) for patients with cervical cancer who underwent radical radiotherapy (RT).
Materials and Methods
Twenty-five patients with cervical cancer were prospectively recruited and treated with external beam RT and brachytherapy. In all patients, except one, chemotherapy was administered concurrently during RT. Whole peripheral blood samples were obtained at least twice from each patient. We performed next-generation sequencing (NGS) for the target-captured libraries (67 oncogenes and human papillomavirus [HPV] type 16/18) using 64 plasma cfDNA samples from the 25 participants. The ratio of HPV cfDNA and the variant allele frequency (VAF) in cfDNA was calculated, and their dynamic changes were monitored. The median follow-up duration was 25.4 months.
Results
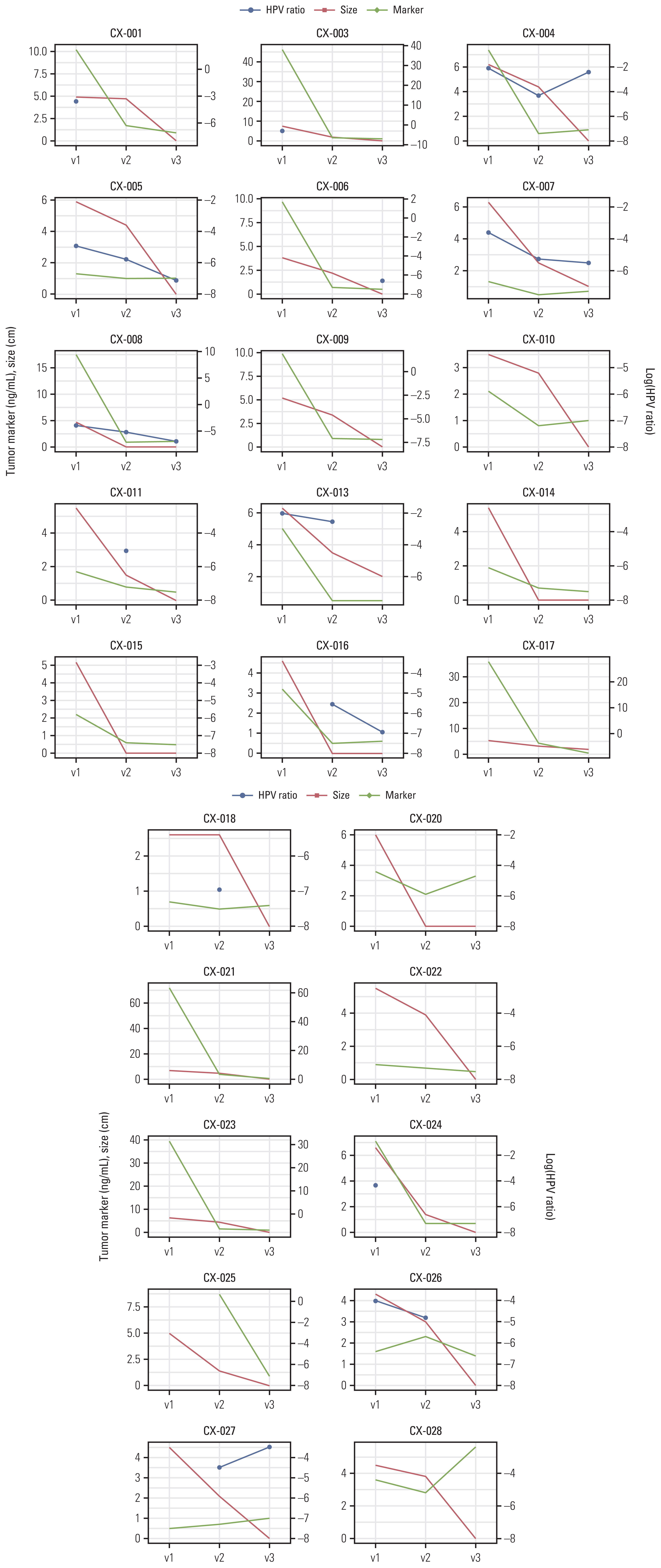
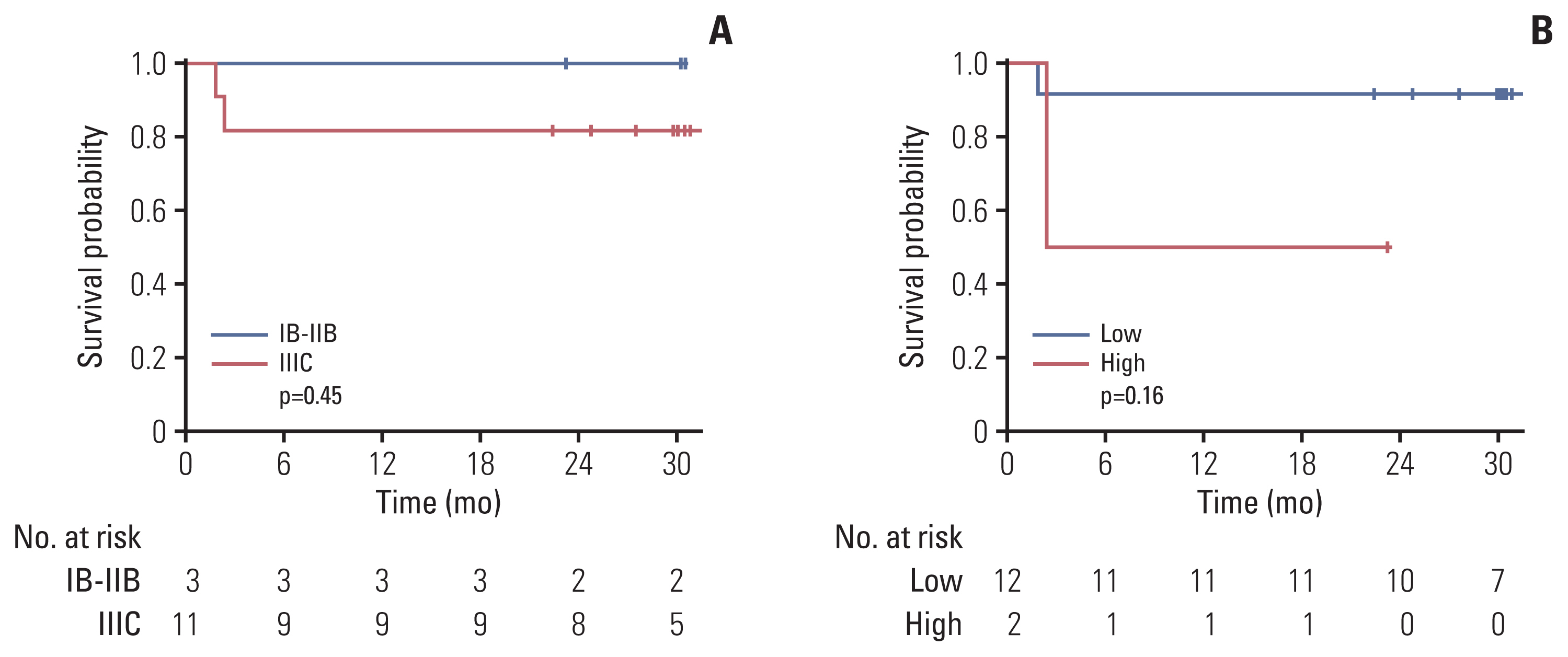
In total, we identified 21,866 cfDNA variants. ARID1A and frameshift variants occupied the largest portion of altered genes and HIGH-grade variant types, respectively. In most cases, tumor shrinkage was followed by a decrease in the HPV ratio; however, an increase in HPV ratio indicated distant metastasis, despite the reduced tumor size. The initial HPV ratio reflecting the tumor burden was likely associated with treatment outcomes (p = 0.16). We did not determine a role for serial changes in the VAF in cfDNA.
Conclusion
Our findings suggest that the HPV cfDNA ratio, calculated after targeted NGS, may be valuable for monitoring and predicting treatment responses. Accordingly, further validation of these findings is warranted.
Keyword
Figure
Reference
-
References
1. Small W Jr, Bacon MA, Bajaj A, Chuang LT, Fisher BJ, Harkenrider MM, et al. Cervical cancer: a global health crisis. Cancer. 2017; 123:2404–12.
Article2. Ferrandina G, Margariti PA, Smaniotto D, Petrillo M, Salerno MG, Fagotti A, et al. Long-term analysis of clinical outcome and complications in locally advanced cervical cancer patients administered concomitant chemoradiation followed by radical surgery. Gynecol Oncol. 2010; 119:404–10.
Article3. Hequet D, Marchand E, Place V, Fourchotte V, De La Rochefordiere A, Dridi S, et al. Evaluation and impact of residual disease in locally advanced cervical cancer after concurrent chemoradiation therapy: results of a multicenter study. Eur J Surg Oncol. 2013; 39:1428–34.
Article4. Kim JY, Byun SJ, Kim YS, Nam JH. Disease courses in patients with residual tumor following concurrent chemoradiotherapy for locally advanced cervical cancer. Gynecol Oncol. 2017; 144:34–9.
Article5. Elit L, Fyles AW, Devries MC, Oliver TK, Fung-Kee-Fung M; Gynecology Cancer Disease Site Group. Follow-up for women after treatment for cervical cancer: a systematic review. Gynecol Oncol. 2009; 114:528–35.
Article6. Lee JH, Lee SW, Kim JR, Kim YS, Yoon MS, Jeong S, et al. Tumour size, volume, and marker expression during radiation therapy can predict survival of cervical cancer patients: a multi-institutional retrospective analysis of KROG 16-01. Gynecol Oncol. 2017; 147:577–84.
Article7. Gadducci A, Tana R, Cosio S, Genazzani AR. The serum assay of tumour markers in the prognostic evaluation, treatment monitoring and follow-up of patients with cervical cancer: a review of the literature. Crit Rev Oncol Hematol. 2008; 66:10–20.
Article8. Moreno-Acosta P, Vallard A, Carrillo S, Gamboa O, Romero-Rojas A, Molano M, et al. Biomarkers of resistance to radiation therapy: a prospective study in cervical carcinoma. Radiat Oncol. 2017; 12:120.
Article9. Ho CK, Kornaga EN, Klimowicz AC, Enwere EK, Dean M, Bebb GD, et al. Expression of DNA damage response proteins in cervical cancer patients treated with radical chemoradiotherapy. Gynecol Oncol. 2017; 145:176–84.
Article10. Chen M, Zhao H. Next-generation sequencing in liquid biopsy: cancer screening and early detection. Hum Genomics. 2019; 13:34.
Article11. Ignatiadis M, Sledge GW, Jeffrey SS. Liquid biopsy enters the clinic: implementation issues and future challenges. Nat Rev Clin Oncol. 2021; 18:297–312.
Article12. Kang Z, Stevanovic S, Hinrichs CS, Cao L. Circulating cell-free DNA for metastatic cervical cancer detection, genotyping, and monitoring. Clin Cancer Res. 2017; 23:6856–62.
Article13. Corcoran RB, Chabner BA. Application of cell-free DNA analysis to cancer treatment. N Engl J Med. 2018; 379:1754–65.
Article14. Mahantshetty U, Teni T, Naga P, Hotwani C, Umesh S, Kannan S, et al. Impact of HPV 16/18 infection on clinical outcomes in locally advanced cervical cancers treated with radical radio (chemo) therapy: a prospective observational study. Gynecol Oncol. 2018; 148:299–304.
Article15. Okuma K, Yamashita H, Yokoyama T, Nakagawa K, Kawana K. Undetected human papillomavirus DNA and uterine cervical carcinoma: association with cancer recurrence. Strahlenther Onkol. 2016; 192:55–62.
Article16. Chung TK, Cheung TH, Yim SF, Yu MY, Chiu RW, Lo KW, et al. Liquid biopsy of PIK3CA mutations in cervical cancer in Hong Kong Chinese women. Gynecol Oncol. 2017; 146:334–9.
Article17. McIntyre JB, Wu JS, Craighead PS, Phan T, Kobel M, Lees-Miller SP, et al. PIK3CA mutational status and overall survival in patients with cervical cancer treated with radical chemoradiotherapy. Gynecol Oncol. 2013; 128:409–14.
Article18. Lee SY, Chae DK, Lee SH, Lim Y, An J, Chae CH, et al. Efficient mutation screening for cervical cancers from circulating tumor DNA in blood. BMC Cancer. 2020; 20:694.
Article19. Mertens F, Johansson B, Hoglund M, Mitelman F. Chromosomal imbalance maps of malignant solid tumors: a cytogenetic survey of 3185 neoplasms. Cancer Res. 1997; 57:2765–80.20. Cho H, Kim JS, Chung H, Perry C, Lee H, Kim JH. Loss of ARID1A/BAF250a expression is linked to tumor progression and adverse prognosis in cervical cancer. Hum Pathol. 2013; 44:1365–74.
Article21. Huang J, Zhao YL, Li Y, Fletcher JA, Xiao S. Genomic and functional evidence for an ARID1A tumor suppressor role. Genes Chromosomes Cancer. 2007; 46:745–50.22. Wiegand KC, Shah SP, Al-Agha OM, Zhao Y, Tse K, Zeng T, et al. ARID1A mutations in endometriosis-associated ovarian carcinomas. N Engl J Med. 2010; 363:1532–43.
Article23. Wang K, Kan J, Yuen ST, Shi ST, Chu KM, Law S, et al. Exome sequencing identifies frequent mutation of ARID1A in molecular subtypes of gastric cancer. Nat Genet. 2011; 43:1219–23.
Article24. Tian J, Geng Y, Lv D, Li P, Cordova M, Liao Y, et al. Using plasma cell-free DNA to monitor the chemoradiotherapy course of cervical cancer. Int J Cancer. 2019; 145:2547–57.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Adjuvant Radiotherapy Following Radical Hysterectomy and Bilateral Pelvic Lymph Node Dissection for the Uterine Cervical Cancer: Prognostic Factors and Failure Patterns
- Stage IB Carcinoma of the Cervix : Prognostic Factors Related to 5-year Disease Free Survival
- Flow Cytometric DNA Content Analysis in uterine Cervical Cancer
- Concurrent Chemoradiotherapy in Cervical Cancer (A New Paradigm in Cervical Cancer Treatment)
- The Detection of Oxygen Free Fadical Scavenger, Superoxide Dismutase(SOD) on the Uterine Cervical Tissue