Cancer Res Treat.
2020 Oct;52(4):1067-1083. 10.4143/crt.2020.187.
Elevated Expression of RIOK1 Is Correlated with Breast Cancer Hormone Receptor Status and Promotes Cancer Progression
- Affiliations
-
- 1Medical School of Nantong University, Nantong, China
- 2Department of Clinical Research Center, Nantong First People’s Hospital, The Second Affiliated Hospital of Nantong University, Nantong, China
- 3Department of General Surgery, Affiliated Hospital of Nantong University, Nantong, China
- 4Research Center of Clinical Medicine, Affiliated Hospital of Nantong University, Nantong, China
- KMID: 2507934
- DOI: http://doi.org/10.4143/crt.2020.187
Abstract
- Purpose
RIOK1 has been proved to play an important role in cancer cell proliferation and migration in various types of cancers—such as colorectal and gastric cancers. However, the expression of RIOK1 in breast cancer (BC) and the relationship between RIOK1 expression and the development of BC are not well characterized. In this study, we assessed the expression of RIOK1 in BC and evaluated the mechanisms underlying its biological function in this disease context.
Materials and Methods
We used immunohistochemistry, western blot and quantitative real-time polymerase chain reaction to evaluate the expression of RIOK1 in BC patients. Then, knockdown or overexpression of RIOK1 were used to evaluate the effect on BC cells in vitro and in vivo. Finally, we predicted miR-204-5p could be a potential regulator of RIOK1.
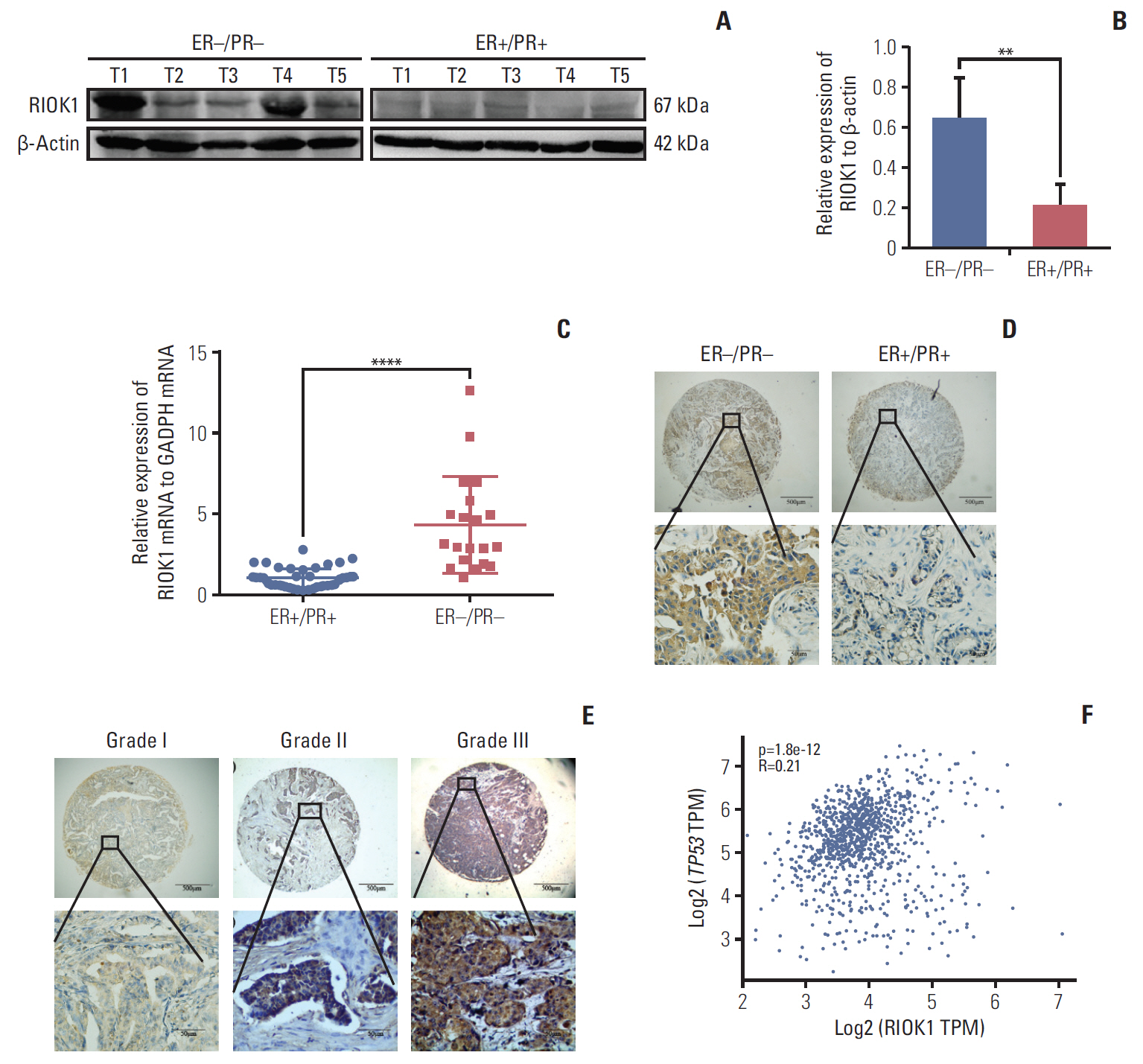
Results
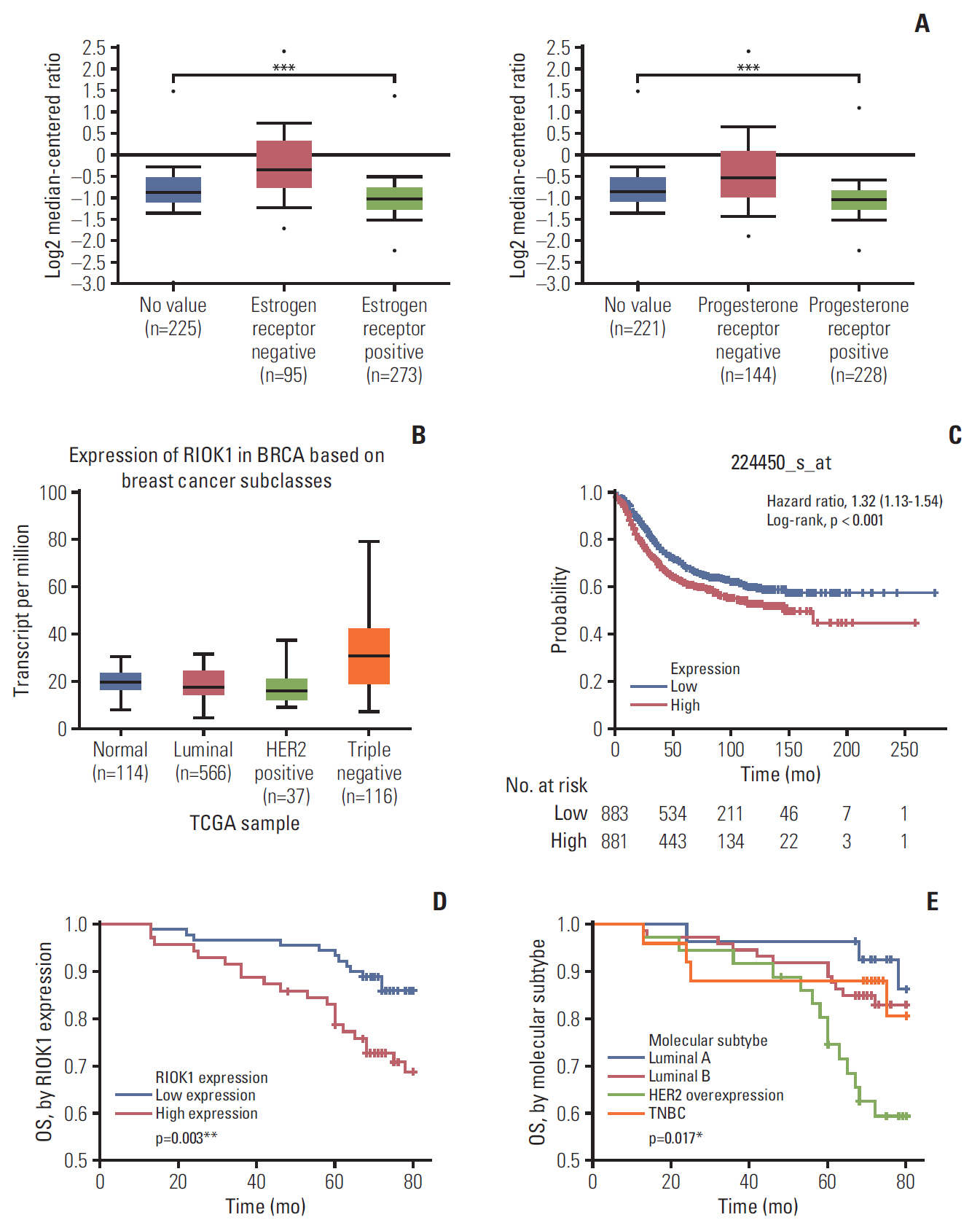
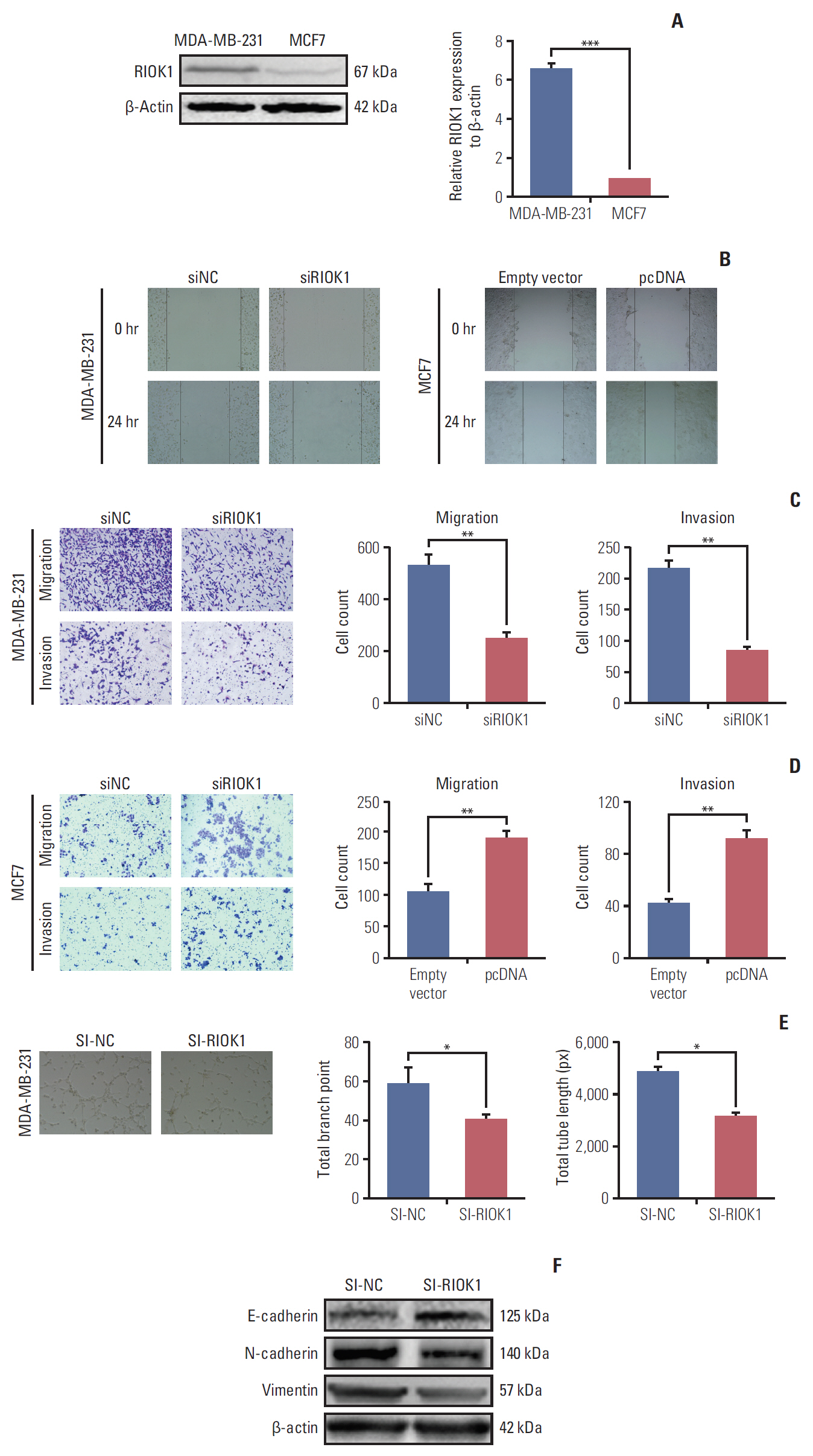
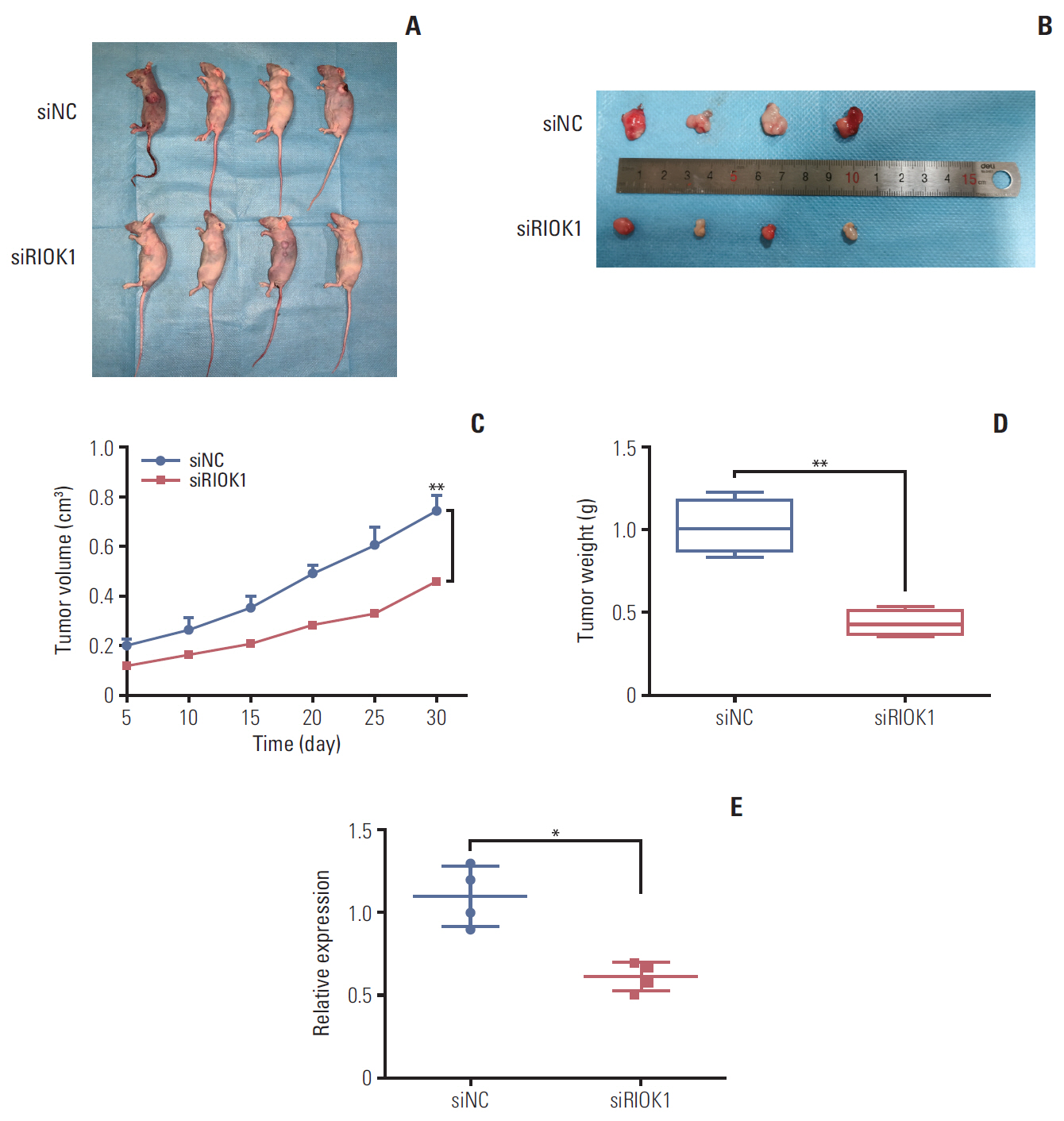
We found that the expression levels of RIOK1 were significantly higher in hormone receptor (HR)–negative BC patients and was associated with tumor grades (p=0.010) and p53 expression (p=0.008) and survival duration (p=0.011). Kaplan-Meier analysis suggested a tendency for the poor prognosis. In vitro, knockdown of RIOK1 could inhibit proliferation, invasion, and induced apoptosis in HR-negative BC cells and inhibited tumorigenesis in vivo, while overexpression of RIOK1 promoted HR-positive tumor progression. MiR-204-5p could regulate RIOK1 expression and be involved in BC progression.
Conclusion
These findings indicate that RIOK1 expression could be a biomarker of HR-negative BC, and it may serve as an effective prognostic indicator and promote BC progression.
Keyword
Figure
Reference
-
References
1. Ferlay J, Soerjomataram I, Dikshit R, Eser S, Mathers C, Rebelo M, et al. Cancer incidence and mortality worldwide: sources, methods and major patterns in GLOBOCAN 2012. Int J Cancer. 2015; 136:E359–86.
Article2. Siegel R, Naishadham D, Jemal A. Cancer statistics, 2012. CA Cancer J Clin. 2012; 62:10–29.
Article3. Coates AS, Winer EP, Goldhirsch A, Gelber RD, Gnant M, Piccart-Gebhart M, et al. Tailoring therapies: improving the management of early breast cancer: St Gallen International Expert Consensus on the Primary Therapy of Early Breast Cancer 2015. Ann Oncol. 2015; 26:1533–46.4. Beith J, Burslem K, Bell R, Woodward N, McCarthy N, De Boer R, et al. Hormone receptor positive, HER2 negative metastatic breast cancer: a systematic review of the current treatment landscape. Asia Pac J Clin Oncol. 2016; 12 Suppl 1:3–18.
Article5. Gerratana L, Fanotto V, Bonotto M, Bolzonello S, Minisini AM, Fasola G, et al. Pattern of metastasis and outcome in patients with breast cancer. Clin Exp Metastasis. 2015; 32:125–33.
Article6. Sihto H, Lundin J, Lundin M, Lehtimaki T, Ristimaki A, Holli K, et al. Breast cancer biological subtypes and protein expression predict for the preferential distant metastasis sites: a nationwide cohort study. Breast Cancer Res. 2011; 13:R87.
Article7. Pienkowski T, Zielinski CC. Trastuzumab treatment in patients with breast cancer and metastatic CNS disease. Ann Oncol. 2010; 21:917–24.8. Vaz-Luis I, Lin NU, Keating NL, Barry WT, Winer EP, Freedman RA. Factors associated with early mortality among patients with de novo metastatic breast cancer: a populationbased study. Oncologist. 2017; 22:386–93.
Article9. LaRonde-LeBlanc N, Wlodawer A. A family portrait of the RIO kinases. J Biol Chem. 2005; 280:37297–300.
Article10. LaRonde NA. The ancient microbial RIO kinases. J Biol Chem. 2014; 289:9488–92.
Article11. Baumas K, Soudet J, Caizergues-Ferrer M, Faubladier M, Henry Y, Mougin A. Human RioK3 is a novel component of cytoplasmic pre-40S pre-ribosomal particles. RNA Biol. 2012; 9:162–74.
Article12. Dez C, Tollervey D. Ribosome synthesis meets the cell cycle. Curr Opin Microbiol. 2004; 7:631–7.
Article13. Kimmelman AC, Hezel AF, Aguirre AJ, Zheng H, Paik JH, Ying H, et al. Genomic alterations link Rho family of GTPases to the highly invasive phenotype of pancreas cancer. Proc Natl Acad Sci U S A. 2008; 105:19372–7.
Article14. Read RD, Fenton TR, Gomez GG, Wykosky J, Vandenberg SR, Babic I, et al. A kinome-wide RNAi screen in Drosophila Glia reveals that the RIO kinases mediate cell proliferation and survival through TORC2-Akt signaling in glioblastoma. PLoS Genet. 2013; 9:e1003253.
Article15. Weinberg F, Reischmann N, Fauth L, Taromi S, Mastroianni J, Kohler M, et al. The atypical kinase RIOK1 promotes tumor growth and invasive behavior. EBioMedicine. 2017; 20:79–97.
Article16. Hong X, Huang H, Qiu X, Ding Z, Feng X, Zhu Y, et al. Targeting posttranslational modifications of RIOK1 inhibits the progression of colorectal and gastric cancers. Elife. 2018; 7:e29511.
Article17. Manning G, Plowman GD, Hunter T, Sudarsanam S. Evolution of protein kinase signaling from yeast to man. Trends Biochem Sci. 2002; 27:514–20.
Article18. Manning G, Whyte DB, Martinez R, Hunter T, Sudarsanam S. The protein kinase complement of the human genome. Science. 2002; 298:1912–34.
Article19. Angermayr M, Roidl A, Bandlow W. Yeast Rio1p is the founding member of a novel subfamily of protein serine kinases involved in the control of cell cycle progression. Mol Microbiol. 2002; 44:309–24.
Article20. Cox AD, Fesik SW, Kimmelman AC, Luo J, Der CJ. Drugging the undruggable RAS: Mission possible? Nat Rev Drug Discov. 2014; 13:828–51.
Article21. Alvarado R, Lari SA, Roses RE, Smith BD, Yang W, Mittendorf EA, et al. Biology, treatment, and outcome in very young and older women with DCIS. Ann Surg Oncol. 2012; 19:3777–84.
Article22. Sionov RV, Haupt Y. The cellular response to p53: the decision between life and death. Oncogene. 1999; 18:6145–57.
Article23. Vousden KH, Lu X. Live or let die: the cell’s response to p53. Nat Rev Cancer. 2002; 2:594–604.
Article24. Kandoth C, McLellan MD, Vandin F, Ye K, Niu B, Lu C, et al. Mutational landscape and significance across 12 major cancer types. Nature. 2013; 502:333–9.
Article25. Langerod A, Zhao H, Borgan O, Nesland JM, Bukholm IR, Ikdahl T, et al. TP53 mutation status and gene expression profiles are powerful prognostic markers of breast cancer. Breast Cancer Res. 2007; 9:R30.
Article26. Miller LD, Smeds J, George J, Vega VB, Vergara L, Ploner A, et al. An expression signature for p53 status in human breast cancer predicts mutation status, transcriptional effects, and patient survival. Proc Natl Acad Sci U S A. 2005; 102:13550–5.27. Darb-Esfahani S, Denkert C, Stenzinger A, Salat C, Sinn B, Schem C, et al. Role of TP53 mutations in triple negative and HER2-positive breast cancer treated with neoadjuvant anthracycline/taxane-based chemotherapy. Oncotarget. 2016; 7:67686–98.28. Li W, Jin X, Zhang Q, Zhang G, Deng X, Ma L. Decreased expression of miR-204 is associated with poor prognosis in patients with breast cancer. Int J Clin Exp Pathol. 2014; 7:3287–92.29. Shen SQ, Huang LS, Xiao XL, Zhu XF, Xiong DD, Cao XM, et al. miR-204 regulates the biological behavior of breast cancer MCF-7 cells by directly targeting FOXA1. Oncol Rep. 2017; 38:368–76.
Article30. Boo L, Ho WY, Mohd Ali N, Yeap SK, Ky H, Chan KG, et al. Phenotypic and microRNA transcriptomic profiling of the MDA-MB-231 spheroid-enriched CSCs with comparison of MCF-7 microRNA profiling dataset. PeerJ. 2017; 5:e3551.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Correlation between COX-2 Expression and Hormone Receptors in Invasive Ductal Breast Cancer
- Hormone Treatment for Breast Cancer
- Leptin and Leptin Receptor Expression in Breast Cancer
- MicroRNA-222 Expression as a Predictive Marker for Tumor Progression in Hormone Receptor-Positive Breast Cancer
- Hormone Therapy for Metastatic Breast Cancer