J Pathol Transl Med.
2019 Sep;53(5):289-297. 10.4132/jptm.2019.06.07.
Clinicopathological Characterization and Prognostic Implication of SMAD4 Expression in Colorectal Carcinoma
- Affiliations
-
- 1Department of Pathology, Seoul National University Hospital, Seoul, Korea. jeongmobae@gmail.com, ghkang@snu.ac.kr
- 2Laboratory of Epigenetics, Cancer Research Institute, Seoul National University College of Medicine, Seoul, Korea.
- KMID: 2459577
- DOI: http://doi.org/10.4132/jptm.2019.06.07
Abstract
- BACKGROUND
SMAD family member 4 (SMAD4) has gained attention as a promising prognostic factor of colorectal cancer (CRC) as well as a key molecule to understand the tumorigenesis and progression of CRC.
METHODS
We retrospectively analyzed 1,281 CRC cases immunohistochemically for their expression status of SMAD4, and correlated this status with clinicopathologic and molecular features of CRCs.
RESULTS
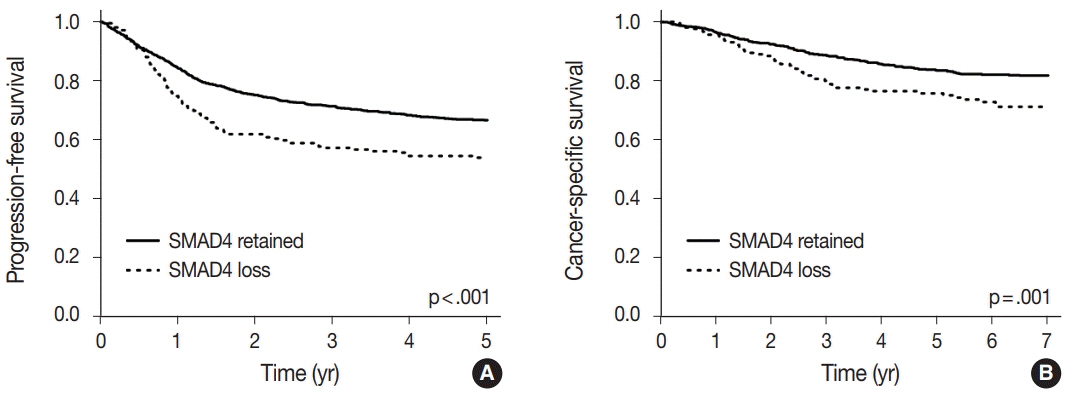
A loss of nuclear SMAD4 was significantly associated with frequent lymphovascular and perineural invasion, tumor budding, fewer tumor-infiltrating lymphocytes, higher pT and pN category, and frequent distant metastasis. In contrast, tumors overexpressing SMAD4 showed a significant association with sporadic microsatellite instability. After adjustment for TNM stage, tumor differentiation, adjuvant chemotherapy, and lymphovascular invasion, the loss of SMAD4 was found to be an independent prognostic factor for worse 5-year progression-free survival (hazard ratio [HR], 1.27; 95% confidence interval [CI], 1.01 to 1.60; p=.042) and 7-year cancer-specific survival (HR, 1.45; 95% CI, 1.06 to 1.99; p=.022).
CONCLUSIONS
We confirmed the value of determining the loss of SMAD4 immunohistochemically as an independent prognostic factor for CRC in general. In addition, we identified some histologic and molecular features that might be clues to elucidate the role of SMAD4 in colorectal tumorigenesis and progression.
Keyword
MeSH Terms
Figure
Reference
-
1. Bray F, Ferlay J, Soerjomataram I, Siegel RL, Torre LA, Jemal A. Global cancer statistics 2018: GLOBOCAN estimates of incidence and mortality worldwide for 36 cancers in 185 countries. CA Cancer J Clin. 2018; 68:394–424.
Article2. Zhao M, Mishra L, Deng CX. The role of TGF-beta/SMAD4 signaling in cancer. Int J Biol Sci. 2018; 14:111–23.3. Schutte M. DPC4/SMAD4 gene alterations in human cancer, and their functional implications. Ann Oncol. 1999; 10 Suppl 4:56–9.
Article4. Du Y, Zhou X, Huang Z, et al. Meta-analysis of the prognostic value of smad4 immunohistochemistry in various cancers. PLoS One. 2014; 9:e110182.
Article5. Wang JD, Jin K, Chen XY, Lv JQ, Ji KW. Clinicopathological significance of SMAD4 loss in pancreatic ductal adenocarcinomas: a systematic review and meta-analysis. Oncotarget. 2017; 8:16704–11.
Article6. Kouvidou C, Latoufis C, Lianou E, et al. Expression of Smad4 and TGF-beta2 in colorectal carcinoma. Anticancer Res. 2006; 26:2901–7.7. Xu WQ, Jiang XC, Zheng L, Yu YY, Tang JM. Expression of TGF-beta1, TbetaRII and Smad4 in colorectal carcinoma. Exp Mol Pathol. 2007; 82:284–91.8. Bacman D, Merkel S, Croner R, Papadopoulos T, Brueckl W, Dimmler A. TGF-beta receptor 2 downregulation in tumour-associated stroma worsens prognosis and high-grade tumours show more tumour-associated macrophages and lower TGF-beta1 expression in colon carcinoma: a retrospective study. BMC Cancer. 2007; 7:156.
Article9. Isaksson-Mettaväinio M, Palmqvist R, Dahlin AM, et al. High SMAD4 levels appear in microsatellite instability and hypermethylated colon cancers, and indicate a better prognosis. Int J Cancer. 2012; 131:779–88.10. Voorneveld PW, Jacobs RJ, De Miranda NF, et al. Evaluation of the prognostic value of pSMAD immunohistochemistry in colorectal cancer. Eur J Cancer Prev. 2013; 22:420–4.
Article11. Kozak MM, von Eyben R, Pai J, et al. Smad4 inactivation predicts for worse prognosis and response to fluorouracil-based treatment in colorectal cancer. J Clin Pathol. 2015; 68:341–5.
Article12. Yan P, Klingbiel D, Saridaki Z, et al. Reduced expression of SMAD4 is associated with poor survival in colon cancer. Clin Cancer Res. 2016; 22:3037–47.
Article13. Jain S, Singhal S, Francis F, et al. Association of overexpression of TIF1gamma with colorectal carcinogenesis and advanced colorectal adenocarcinoma. World J Gastroenterol. 2011; 17:3994–4000.14. Handra-Luca A, Olschwang S, Fléjou JF. SMAD4 protein expression and cell proliferation in colorectal adenocarcinomas. Virchows Arch. 2011; 459:511–9.
Article15. Chung Y, Wi YC, Kim Y, et al. The Smad4/PTEN expression pattern predicts clinical outcomes in colorectal adenocarcinoma. J Pathol Transl Med. 2018; 52:37–44.
Article16. Oyanagi H, Shimada Y, Nagahashi M, et al. SMAD4 alteration associates with invasive-front pathological markers and poor prognosis in colorectal cancer. Histopathology. 2019; 74:873–82.17. Wasserman I, Lee LH, Ogino S, et al. SMAD4 loss in colorectal cancer patients correlates with recurrence, loss of immune infiltrate, and chemoresistance. Clin Cancer Res. 2019; 25:1948–56.
Article18. Alhopuro P, Alazzouzi H, Sammalkorpi H, et al. SMAD4 levels and response to 5-fluorouracil in colorectal cancer. Clin Cancer Res. 2005; 11:6311–6.
Article19. Boulay JL, Mild G, Lowy A, et al. SMAD4 is a predictive marker for 5-fluorouracil-based chemotherapy in patients with colorectal cancer. Br J Cancer. 2002; 87:630–4.
Article20. Alazzouzi H, Alhopuro P, Salovaara R, et al. SMAD4 as a prognostic marker in colorectal cancer. Clin Cancer Res. 2005; 11:2606–11.
Article21. Isaksson-Mettavainio M, Palmqvist R, Forssell J, Stenling R, Oberg A. SMAD4/DPC4 expression and prognosis in human colorectal cancer. Anticancer Res. 2006; 26:507–10.22. Mesker WE, Liefers GJ, Junggeburt JM, et al. Presence of a high amount of stroma and downregulation of SMAD4 predict for worse survival for stage I-II colon cancer patients. Cell Oncol. 2009; 31:169–78.
Article23. Baraniskin A, Munding J, Schulmann K, et al. Prognostic value of reduced SMAD4 expression in patients with metastatic colorectal cancer under oxaliplatin-containing chemotherapy: a translational study of the AIO colorectal study group. Clin Colorectal Cancer. 2011; 10:24–9.
Article24. Wang C, Zhou Y, Ruan R, Zheng M, Han W, Liao L. High expression of COUP-TF II cooperated with negative Smad4 expression predicts poor prognosis in patients with colorectal cancer. Int J Clin Exp Pathol. 2015; 8:7112–21.25. Coates RF, Gardner JA, Gao Y, et al. Significance of positive and inhibitory regulators in the TGF-beta signaling pathway in colorectal cancers. Hum Pathol. 2017; 66:34–9.26. Xie W, Rimm DL, Lin Y, Shih WJ, Reiss M. Loss of Smad signaling in human colorectal cancer is associated with advanced disease and poor prognosis. Cancer J. 2003; 9:302–12.
Article27. Gulubova M, Manolova I, Ananiev J, Julianov A, Yovchev Y, Peeva K. Role of TGF-beta1, its receptor TGFbetaRII, and Smad proteins in the progression of colorectal cancer. Int J Colorectal Dis. 2010; 25:591–9.28. Chun HK, Jung KU, Choi YL, et al. Low expression of transforming growth factor beta-1 in cancer tissue predicts a poor prognosis for patients with stage III rectal cancers. Oncology. 2014; 86:159–69.
Article29. Voorneveld PW, Jacobs RJ, Kodach LL, Hardwick JC. A Meta-analysis of SMAD4 immunohistochemistry as a prognostic marker in colorectal cancer. Transl Oncol. 2015; 8:18–24.
Article30. Zhang B, Zhang B, Chen X, et al. Loss of Smad4 in colorectal cancer induces resistance to 5-fluorouracil through activating Akt pathway. Br J Cancer. 2014; 110:946–57.
Article31. Voorneveld PW, Kodach LL, Jacobs RJ, et al. The BMP pathway either enhances or inhibits the Wnt pathway depending on the SMAD4 and p53 status in CRC. Br J Cancer. 2015; 112:122–30.32. Chow E, Macrae F. A review of juvenile polyposis syndrome. J Gastroenterol Hepatol. 2005; 20:1634–40.
Article33. Ahn BK, Jang SH, Paik SS, Lee KH. Smad4 may help to identify a subset of colorectal cancer patients with early recurrence after curative therapy. Hepatogastroenterology. 2011; 58:1933–6.
Article34. Bae JM, Kim JH, Kwak Y, et al. Distinct clinical outcomes of two CIMP-positive colorectal cancer subtypes based on a revised CIMP classification system. Br J Cancer. 2017; 116:1012–20.
Article35. Bankhead P, Loughrey MB, Fernández JA, et al. QuPath: open source software for digital pathology image analysis. Sci Rep. 2017; 7:16878.
Article36. Bae JM, Lee TH, Cho NY, Kim TY, Kang GH. Loss of CDX2 expression is associated with poor prognosis in colorectal cancer patients. World J Gastroenterol. 2015; 21:1457–67.
Article37. Boland CR, Thibodeau SN, Hamilton SR, et al. A National Cancer Institute Workshop on Microsatellite Instability for cancer detection and familial predisposition: development of international criteria for the determination of microsatellite instability in colorectal cancer. Cancer Res. 1998; 58:5248–57.38. Weisenberger DJ, Siegmund KD, Campan M, et al. CpG island methylator phenotype underlies sporadic microsatellite instability and is tightly associated with BRAF mutation in colorectal cancer. Nat Genet. 2006; 38:787–93.39. Jung B, Staudacher JJ, Beauchamp D. Transforming growth factor beta superfamily signaling in development of colorectal cancer. Gastroenterology. 2017; 152:36–52.40. Salovaara R, Roth S, Loukola A, et al. Frequent loss of SMAD4/DPC4 protein in colorectal cancers. Gut. 2002; 51:56–9.
Article41. Ilyas M, Efstathiou JA, Straub J, Kim HC, Bodmer WF. Transforming growth factor beta stimulation of colorectal cancer cell lines: type II receptor bypass and changes in adhesion molecule expression. Proc Natl Acad Sci U S A. 1999; 96:3087–91.42. Deacu E, Mori Y, Sato F, et al. Activin type II receptor restoration in ACVR2-deficient colon cancer cells induces transforming growth factor-beta response pathway genes. Cancer Res. 2004; 64:7690–6.43. Baker K, Raut P, Jass JR. Microsatellite unstable colorectal cancer cell lines with truncating TGFbetaRII mutations remain sensitive to endogenous TGFbeta. J Pathol. 2007; 213:257–65.44. de Miranda NF, van Dinther M, van den Akker BE, van Wezel T, ten Dijke P, Morreau H. Transforming growth factor beta signaling in colorectal cancer cells with microsatellite instability despite biallelic mutations in TGFBR2. Gastroenterology. 2015; 148:1427–37. e8.45. Barros R, Pereira B, Duluc I, et al. Key elements of the BMP/SMAD pathway co-localize with CDX2 in intestinal metaplasia and regulate CDX2 expression in human gastric cell lines. J Pathol. 2008; 215:411–20.
Article46. Barros R, Mendes N, Howe JR, et al. Juvenile polyps have gastric differentiation with MUC5AC expression and downregulation of CDX2 and SMAD4. Histochem Cell Biol. 2009; 131:765–72.
Article47. Savagner P. The epithelial-mesenchymal transition (EMT) phenomenon. Ann Oncol. 2010; 21 Suppl 7:vii89–92.
Article48. Ioannou M, Kouvaras E, Papamichali R, Samara M, Chiotoglou I, Koukoulis G. Smad4 and epithelial-mesenchymal transition proteins in colorectal carcinoma: an immunohistochemical study. J Mol Histol. 2018; 49:235–44.
Article49. Deckers M, van Dinther M, Buijs J, et al. The tumor suppressor Smad4 is required for transforming growth factor beta-induced epithelial to mesenchymal transition and bone metastasis of breast cancer cells. Cancer Res. 2006; 66:2202–9.50. Itatani Y, Kawada K, Fujishita T, et al. Loss of SMAD4 from colorectal cancer cells promotes CCL15 expression to recruit CCR1+ myeloid cells and facilitate liver metastasis. Gastroenterology. 2013; 145:1064–75. e11.
Article51. Inamoto S, Itatani Y, Yamamoto T, et al. Loss of SMAD4 promotes colorectal cancer progression by accumulation of myeloid-derived suppressor cells through the CCL15-CCR1 chemokine axis. Clin Cancer Res. 2016; 22:492–501.
Article52. Yamamoto T, Kawada K, Itatani Y, et al. Loss of SMAD4 promotes lung metastasis of colorectal cancer by accumulation of CCR1+ tumor-associated neutrophils through CCL15-CCR1 axis. Clin Cancer Res. 2017; 23:833–44.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Clinicopathological Significance of SMAD4 Expression in Breast Cancer
- The Smad4/PTEN Expression Pattern Predicts Clinical Outcomes in Colorectal Adenocarcinoma
- Correlation between Expression of c-erbB-2 Oncogene and Various Prognostic Factors in the Colorectal Carcinoma
- Smad4 Expression in Gastric Adenocarcinoma
- Inhibitors of Apoptosis Proteins Expression and Their Prognostic Significance in Colorectal Carcinoma