Allergy Asthma Immunol Res.
2019 Sep;11(5):691-708. 10.4168/aair.2019.11.5.691.
In-Depth, Proteomic Analysis of Nasal Secretions from Patients With Chronic Rhinosinusitis and Nasal Polyps
- Affiliations
-
- 1Obstructive Upper airway Research (OUaR) Laboratory, Department of Pharmacology, Seoul National University College of Medicine, Seoul, Korea. charlie@snu.ac.kr
- 2Department of Biomedical Sciences, Seoul National University Graduate School, Seoul, Korea.
- 3Proteomics core facility, Biomedical Research Institute, Seoul National University Hospital, Seoul, Korea.
- 4Department of Otorhinolaryngology, Armed Forces Capital Hospital, Seongnam, Korea.
- 5Department of Otorhinolaryngology-Head and Neck Surgery, Boramae Medical Center, Seoul, Korea.
- 6Department of Otorhinolaryngology-Head and Neck Surgery, Chungnam National University Hospital, Daejeon, Korea.
- 7Department of Otorhinolaryngology-Head and Neck Surgery, Dankook University Hospital, Cheonan, Korea.
- 8Department of Otorhinolaryngology-Head and Neck Surgery, Hallym University Hospital, Pyongchon, Korea.
- 9Ischemic/hypoxic disease institute, Seoul National University College of Medicine, Seoul, Korea.
- 10Cancer Research Institute, Seoul National University College of Medicine, Seoul, Korea.
- 11Department of Otorhinolaryngology-Head and Neck Surgery, Seoul National University Hospital, Seoul, Korea.
- 12Clinical Mucosal Immunology Study Group, Seoul, Korea.
- KMID: 2452759
- DOI: http://doi.org/10.4168/aair.2019.11.5.691
Abstract
- PURPOSE
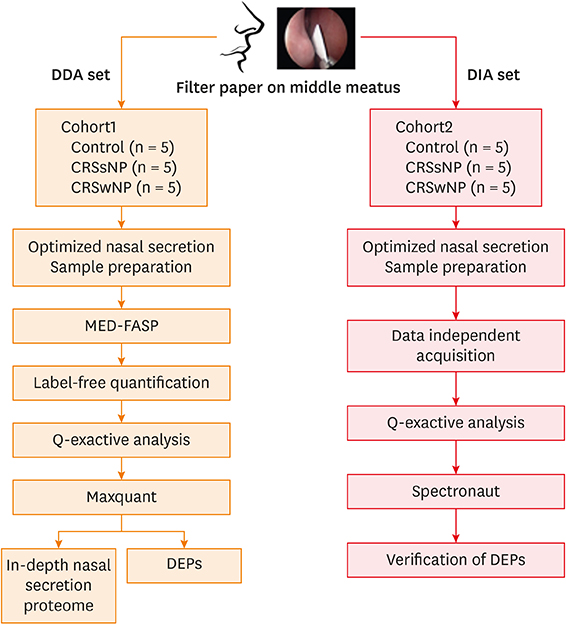
Chronic rhinosinusitis (CRS) is a complex immunological condition, and novel experimental modalities are required to explore various clinical and pathophysiological endotypes; mere evaluation of nasal polyp (NP) status is inadequate. Therefore, we collected patient nasal secretions on filter paper and characterized the proteomes.
METHODS
We performed liquid chromatography-mass spectrometry (MS)/MS in the data-dependent acquisition (DDA) and data-independent acquisition (DIA) modes. Nasal secretions were collected from 10 controls, 10 CRS without NPs (CRSsNP) and 10 CRS with NPs (CRSwNP). We performed Orbitrap MS-based proteomic analysis in the DDA (5 controls, 5 CRSsNP and 5 CRSwNP) and the DIA (5 controls, 5 CRSsNP and 5 CRSwNP) modes, followed by a statistical analysis and a hierarchical clustering to identify differentially expressed proteins in the 3 groups.
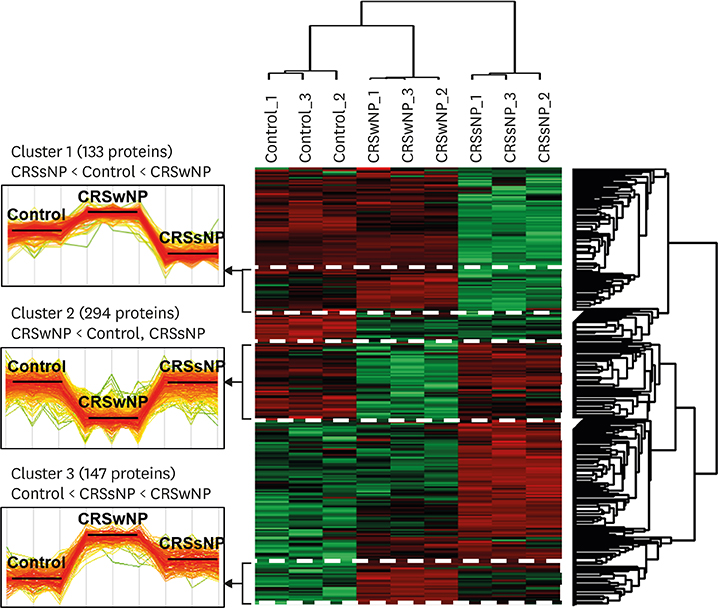
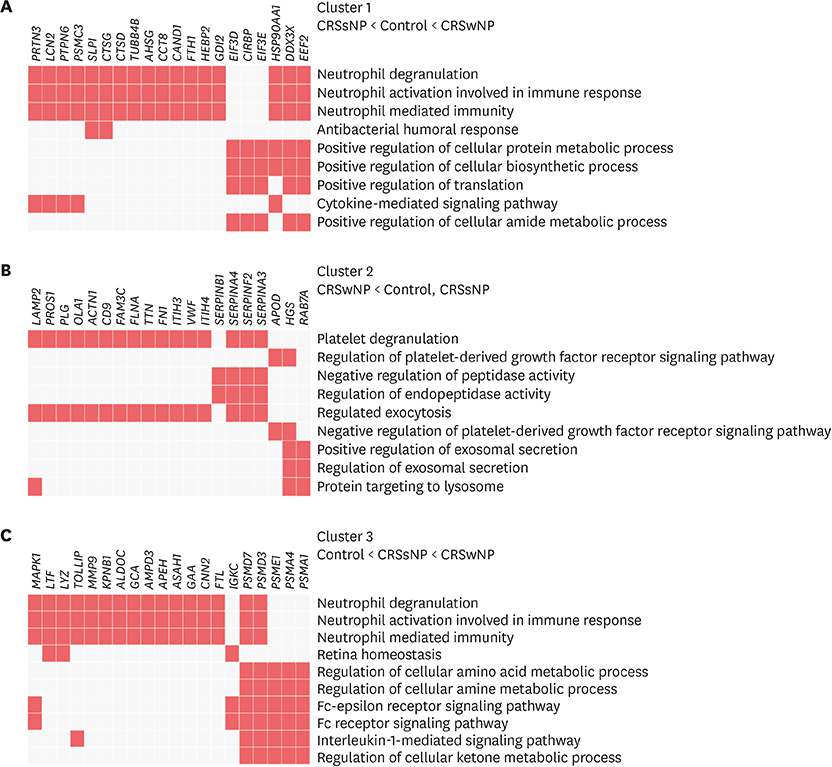
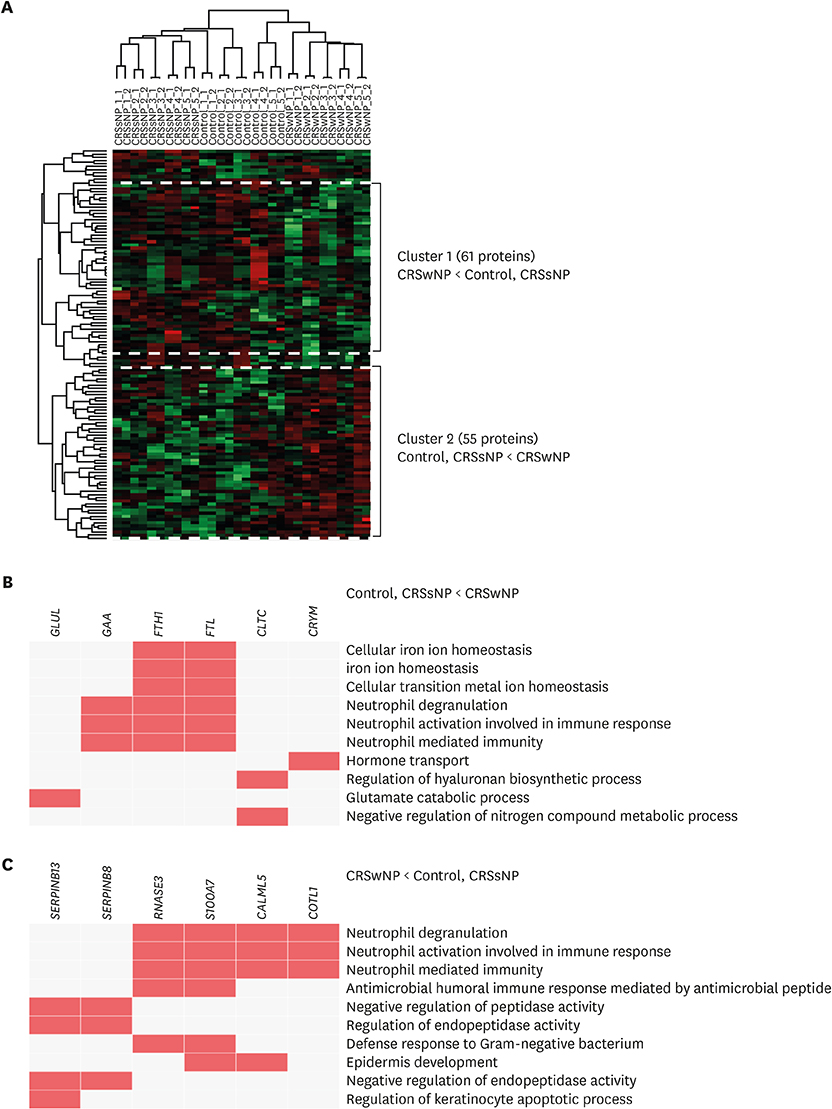
RESULTS
We identified 2,020 proteins in nasal secretions. Canonical pathway analysis and gene ontology (GO) evaluation revealed that interleukin (IL)-7, IL-9, IL-17A and IL-22 signaling and neutrophil-mediated immune responses like neutrophil degranulation and activation were significantly increased in CRSwNP compared to control. The GO terms related to the iron ion metabolism that may be associated with CRS and NP development.
CONCLUSIONS
Collection of nasal secretions on the filter paper is a practical and non-invasive method for in-depth study of nasal proteomics. Our proteomic signatures also support that Asian NPs could be characterized as non-eosinophilic inflammation features. Therefore, the proteomic profiling of nasal secretions from CRS patients may enhance our understanding of CRS endotypes.
Keyword
MeSH Terms
Figure
Cited by 1 articles
-
Evaluation of Neo-Osteogenesis in Eosinophilic Chronic Rhinosinusitis Using a Nasal Polyp Murine Model
Roza Khalmuratova, Mingyu Lee, Jong-Wan Park, Hyun-Woo Shin
Allergy Asthma Immunol Res. 2020;12(2):306-321. doi: 10.4168/aair.2020.12.2.306.
Reference
-
1. Fokkens WJ, Lund VJ, Mullol J, Bachert C, Alobid I, Baroody F, et al. EPOS 2012: European position paper on rhinosinusitis and nasal polyps 2012. A summary for otorhinolaryngologists. Rhinology. 2012; 50:1–12.
Article2. Rudmik L. Economics of chronic rhinosinusitis. Curr Allergy Asthma Rep. 2017; 17:20.
Article3. Bachert C, Zhang N, Hellings PW, Bousquet J. Endotype-driven care pathways in patients with chronic rhinosinusitis. J Allergy Clin Immunol. 2018; 141:1543–1551.
Article4. Kim DW, Cho SH. Emerging endotypes of chronic rhinosinusitis and its application to precision medicine. Allergy Asthma Immunol Res. 2017; 9:299–306.
Article5. Liao B, Liu JX, Li ZY, Zhen Z, Cao PP, Yao Y, et al. Multidimensional endotypes of chronic rhinosinusitis and their association with treatment outcomes. Allergy. 2018; 73:1459–1469.
Article6. Tomassen P, Vandeplas G, Van Zele T, Cardell LO, Arebro J, Olze H, et al. Inflammatory endotypes of chronic rhinosinusitis based on cluster analysis of biomarkers. J Allergy Clin Immunol. 2016; 137:1449–1456.e4.
Article7. Mörtstedt H, Ali N, Kåredal M, Jacobsson H, Rietz E, Diab KK, et al. Targeted proteomic analyses of nasal lavage fluid in persulfate-challenged hairdressers with bleaching powder-associated rhinitis. J Proteome Res. 2015; 14:860–873.
Article8. Biswas K, Wagner Mackenzie B, Waldvogel-Thurlow S, Middleditch M, Jullig M, Zoing M, et al. Differentially regulated host proteins associated with chronic rhinosinusitis are correlated with the sinonasal microbiome. Front Cell Infect Microbiol. 2017; 7:504.
Article9. Tomazic PV, Birner-Gruenberger R, Leitner A, Spoerk S, Lang-Loidolt D. Seasonal proteome changes of nasal mucus reflect perennial inflammatory response and reduced defence mechanisms and plasticity in allergic rhinitis. J Proteomics. 2016; 133:153–160.
Article10. Tomazic PV, Birner-Gruenberger R, Leitner A, Obrist B, Spoerk S, Lang-Loidolt D. Nasal mucus proteomic changes reflect altered immune responses and epithelial permeability in patients with allergic rhinitis. J Allergy Clin Immunol. 2014; 133:741–750.
Article11. Csősz É, Kalló G, Márkus B, Deák E, Csutak A, Tőzsér J. Quantitative body fluid proteomics in medicine - a focus on minimal invasiveness. J Proteomics. 2017; 153:30–43.
Article12. Krismer B, Weidenmaier C, Zipperer A, Peschel A. The commensal lifestyle of Staphylococcus aureus and its interactions with the nasal microbiota. Nat Rev Microbiol. 2017; 15:675–687.13. Cole AM, Liao HI, Stuchlik O, Tilan J, Pohl J, Ganz T. Cationic polypeptides are required for antibacterial activity of human airway fluid. J Immunol. 2002; 169:6985–6991.
Article14. Kirkeby L, Rasmussen TT, Reinholdt J, Kilian M. Immunoglobulins in nasal secretions of healthy humans: structural integrity of secretory immunoglobulin A1 (IgA1) and occurrence of neutralizing antibodies to IgA1 proteases of nasal bacteria. Clin Diagn Lab Immunol. 2000; 7:31–39.
Article15. Tewfik MA, Latterich M, DiFalco MR, Samaha M. Proteomics of nasal mucus in chronic rhinosinusitis. Am J Rhinol. 2007; 21:680–685.
Article16. Alam R, Sim TC, Hilsmeier K, Grant JA. Development of a new technique for recovery of cytokines from inflammatory sites in situ . J Immunol Methods. 1992; 155:25–29.17. Badorrek P, Müller M, Koch W, Hohlfeld JM, Krug N. Specificity and reproducibility of nasal biomarkers in patients with allergic rhinitis after allergen challenge chamber exposure. Ann Allergy Asthma Immunol. 2017; 118:290–297.
Article18. Kim MS, Pinto SM, Getnet D, Nirujogi RS, Manda SS, Chaerkady R, et al. A draft map of the human proteome. Nature. 2014; 509:575–581.19. Doerr A. DIA mass spectrometry. Nat Methods. 2015; 12:35.
Article20. Kulak NA, Pichler G, Paron I, Nagaraj N, Mann M. Minimal, encapsulated proteomic-sample processing applied to copy-number estimation in eukaryotic cells. Nat Methods. 2014; 11:319–324.
Article21. Hu A, Noble WS, Wolf-Yadlin A. Technical advances in proteomics: new developments in data-independent acquisition. F1000 Res. 2016; 5:F1000.
Article22. Gillet LC, Navarro P, Tate S, Röst H, Selevsek N, Reiter L, et al. Targeted data extraction of the MS/MS spectra generated by data-independent acquisition: a new concept for consistent and accurate proteome analysis. Mol Cell Proteomics. 2012; 11:O111.016717.
Article23. Wiśniewski JR, Gaugaz FZ. Fast and sensitive total protein and Peptide assays for proteomic analysis. Anal Chem. 2015; 87:4110–4116.
Article24. Han D, Jin J, Woo J, Min H, Kim Y. Proteomic analysis of mouse astrocytes and their secretome by a combination of FASP and StageTip-based, high pH, reversed-phase fractionation. Proteomics. 2014; 14:1604–1609.
Article25. Woo J, Han D, Park J, Kim SJ, Kim Y. In-depth characterization of the secretome of mouse CNS cell lines by LC-MS/MS without prefractionation. Proteomics. 2015; 15:3617–3622.
Article26. Lee H, Kim K, Woo J, Park J, Kim H, Lee KE, et al. Quantitative proteomic analysis identifies AHNAK (neuroblast differentiation-associated protein AHNAK) as a novel candidate biomarker for bladder urothelial carcinoma diagnosis by liquid-based cytology. Mol Cell Proteomics. 2018; 17:1788–1802.
Article27. Bruderer R, Bernhardt OM, Gandhi T, Miladinović SM, Cheng LY, Messner S, et al. Extending the limits of quantitative proteome profiling with data-independent acquisition and application to acetaminophen-treated three-dimensional liver microtissues. Mol Cell Proteomics. 2015; 14:1400–1410.
Article28. Cox J, Neuhauser N, Michalski A, Scheltema RA, Olsen JV, Mann M. Andromeda: a peptide search engine integrated into the MaxQuant environment. J Proteome Res. 2011; 10:1794–1805.
Article29. Ndika J, Airaksinen L, Suojalehto H, Karisola P, Fyhrquist N, Puustinen A, et al. Epithelial proteome profiling suggests the essential role of interferon-inducible proteins in patients with allergic rhinitis. J Allergy Clin Immunol. 2017; 140:1288–1298.
Article30. Cox J, Hein MY, Luber CA, Paron I, Nagaraj N, Mann M. Accurate proteome-wide label-free quantification by delayed normalization and maximal peptide ratio extraction, termed MaxLFQ. Mol Cell Proteomics. 2014; 13:2513–2526.
Article31. Reiter L, Rinner O, Picotti P, Hüttenhain R, Beck M, Brusniak MY, et al. mProphet: automated data processing and statistical validation for large-scale SRM experiments. Nat Methods. 2011; 8:430–435.
Article32. Tyanova S, Temu T, Sinitcyn P, Carlson A, Hein MY, Geiger T, et al. The Perseus computational platform for comprehensive analysis of (prote)omics data. Nat Methods. 2016; 13:731–740.
Article33. Deeb SJ, Tyanova S, Hummel M, Schmidt-Supprian M, Cox J, Mann M. Machine learning-based classification of diffuse large B-cell lymphoma patients by their protein expression profiles. Mol Cell Proteomics. 2015; 14:2947–2960.
Article34. Perez-Riverol Y, Csordas A, Bai J, Bernal-Llinares M, Hewapathirana S, Kundu DJ, et al. The PRIDE database and related tools and resources in 2019: improving support for quantification data. Nucleic Acids Res. 2019; 47:D442–50.
Article35. Mukaka MM. Statistics corner: a guide to appropriate use of correlation coefficient in medical research. Malawi Med J. 2012; 24:69–71.36. Kuleshov MV, Jones MR, Rouillard AD, Fernandez NF, Duan Q, Wang Z, et al. Enrichr: a comprehensive gene set enrichment analysis web server 2016 update. Nucleic Acids Res. 2016; 44:W90-7.
Article37. Arosio P, Ingrassia R, Cavadini P. Ferritins: a family of molecules for iron storage, antioxidation and more. Biochim Biophys Acta. 2009; 1790:589–599.
Article38. Zandman-Goddard G, Shoenfeld Y. Ferritin in autoimmune diseases. Autoimmun Rev. 2007; 6:457–463.
Article39. Shi LL, Xiong P, Zhang L, Cao PP, Liao B, Lu X, et al. Features of airway remodeling in different types of Chinese chronic rhinosinusitis are associated with inflammation patterns. Allergy. 2013; 68:101–109.
Article40. Zheng Y, Niyonsaba F, Ushio H, Ikeda S, Nagaoka I, Okumura K, et al. Microbicidal protein psoriasin is a multifunctional modulator of neutrophil activation. Immunology. 2008; 124:357–367.
Article41. Kim DK, Eun KM, Kim MK, Cho D, Han SA, Han SY, et al. Comparison between signature cytokines of nasal tissues in subtypes of chronic rhinosinusitis. Allergy Asthma Immunol Res. 2019; 11:201–211.
Article42. Cao PP, Li HB, Wang BF, Wang SB, You XJ, Cui YH, et al. Distinct immunopathologic characteristics of various types of chronic rhinosinusitis in adult Chinese. J Allergy Clin Immunol. 2009; 124:478–484.
Article43. Lee M, Kim DW, Yoon H, So D, Khalmuratova R, Rhee CS, et al. Sirtuin 1 attenuates nasal polypogenesis by suppressing epithelial-to-mesenchymal transition. J Allergy Clin Immunol. 2016; 137:87–98.e7.
Article44. Nograles KE, Zaba LC, Guttman-Yassky E, Fuentes-Duculan J, Suárez-Fariñas M, Cardinale I, et al. Th17 cytokines interleukin (IL)-17 and IL-22 modulate distinct inflammatory and keratinocyte-response pathways. Br J Dermatol. 2008; 159:1092–1102.
Article45. Kotz KT, Xiao W, Miller-Graziano C, Qian WJ, Russom A, Warner EA, et al. Clinical microfluidics for neutrophil genomics and proteomics. Nat Med. 2010; 16:1042–1047.
Article46. Tieu DD, Kern RC, Schleimer RP. Alterations in epithelial barrier function and host defense responses in chronic rhinosinusitis. J Allergy Clin Immunol. 2009; 124:37–42.
Article47. Luo Q, Zhang J, Wang H, Chen F, Luo X, Miao B, et al. Expression and regulation of transcription factor FoxA2 in chronic rhinosinusitis with and without nasal polyps. Allergy Asthma Immunol Res. 2015; 7:458–466.
Article48. Khalmuratova R, Park JW, Shin HW. Immune cell responses and mucosal barrier disruptions in chronic rhinosinusitis. Immune Netw. 2017; 17:60–67.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Medical treatment according to phenotypes of chronic rhinosinusitis
- Practical Review of Biologics in Chronic Rhinosinusitis With Nasal Polyps
- Tissue Remodeling in Rhinosinusitis
- The Role of Superantigen in Nasal Polypogenesis
- Role of Fungal and Bacterial Superantigen in the Pathogenesis of Chronic Rhinosinusitis with Polyps