Cancer Res Treat.
2022 Apr;54(2):597-612. 10.4143/crt.2021.752.
Plasma Circulating Tumor DNA in Patients with Primary Central Nervous System Lymphoma
- Affiliations
-
- 1Division of Hematology-oncology, Department of Medicine, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
- 2Samsung Genome Institute, Samsung Medical Center, Seoul, Korea
- 3Department of Health Sciences and Technology, Samsung Advanced Institute of Health Science and Technology, Sungkyunkwan University, Seoul, Korea
- 4GENINUS Inc., Seoul, Korea
- 5Department of Pathology, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
- 6Department of Molecular Cell Biology, Sungkyunkwan University School of Medicine, Suwon, Korea
- 7Division of Hematology-Oncology, Department of Internal Medicine, Ewha Medical Research Center, School of Medicine, Ewha Womans University, Seoul, Korea
- 8Department of Laboratory Medicine and Genetics, Samsung Medical Center, Sungkyunkwan University School of Medicine, Seoul, Korea
- KMID: 2528229
- DOI: http://doi.org/10.4143/crt.2021.752
Abstract
- Purpose
Analysis of circulating tumor DNA (ctDNA) in blood could allow noninvasive genetic analysis of primary tumors. Although there have been unmet needs for noninvasive methods in patients with primary central nervous system lymphoma (PCNSL), it is still not determined whether plasma ctDNA analysis could be useful for patients with PCNSL.
Materials and Methods
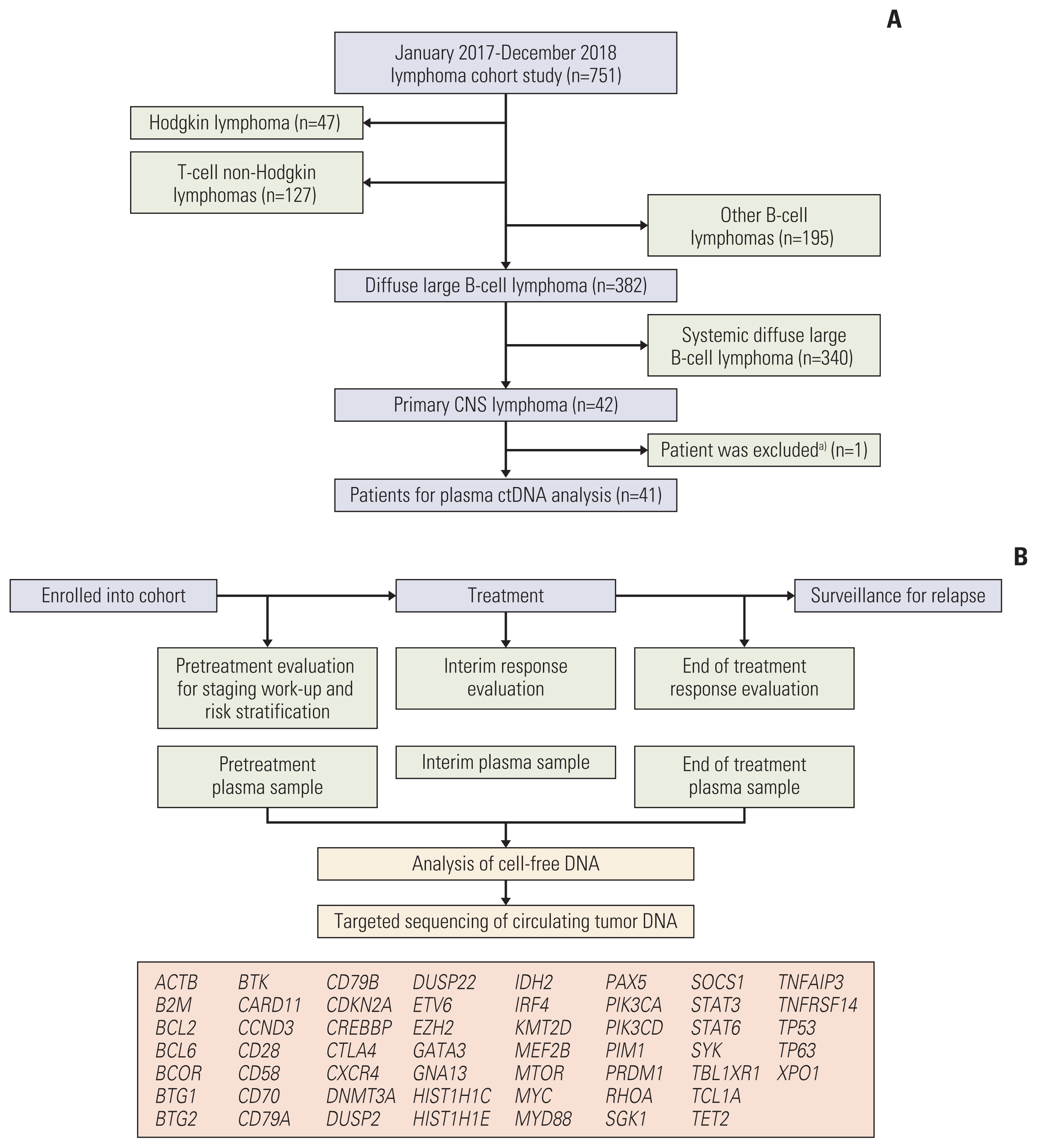
Targeted deep sequencing of 54 genes was performed in cell-free DNA isolated from plasma samples collected pretreatment, during treatment, and at the end of treatment in 42 consecutively diagnosed PCNSL patients between January 2017 and December 2018.
Results
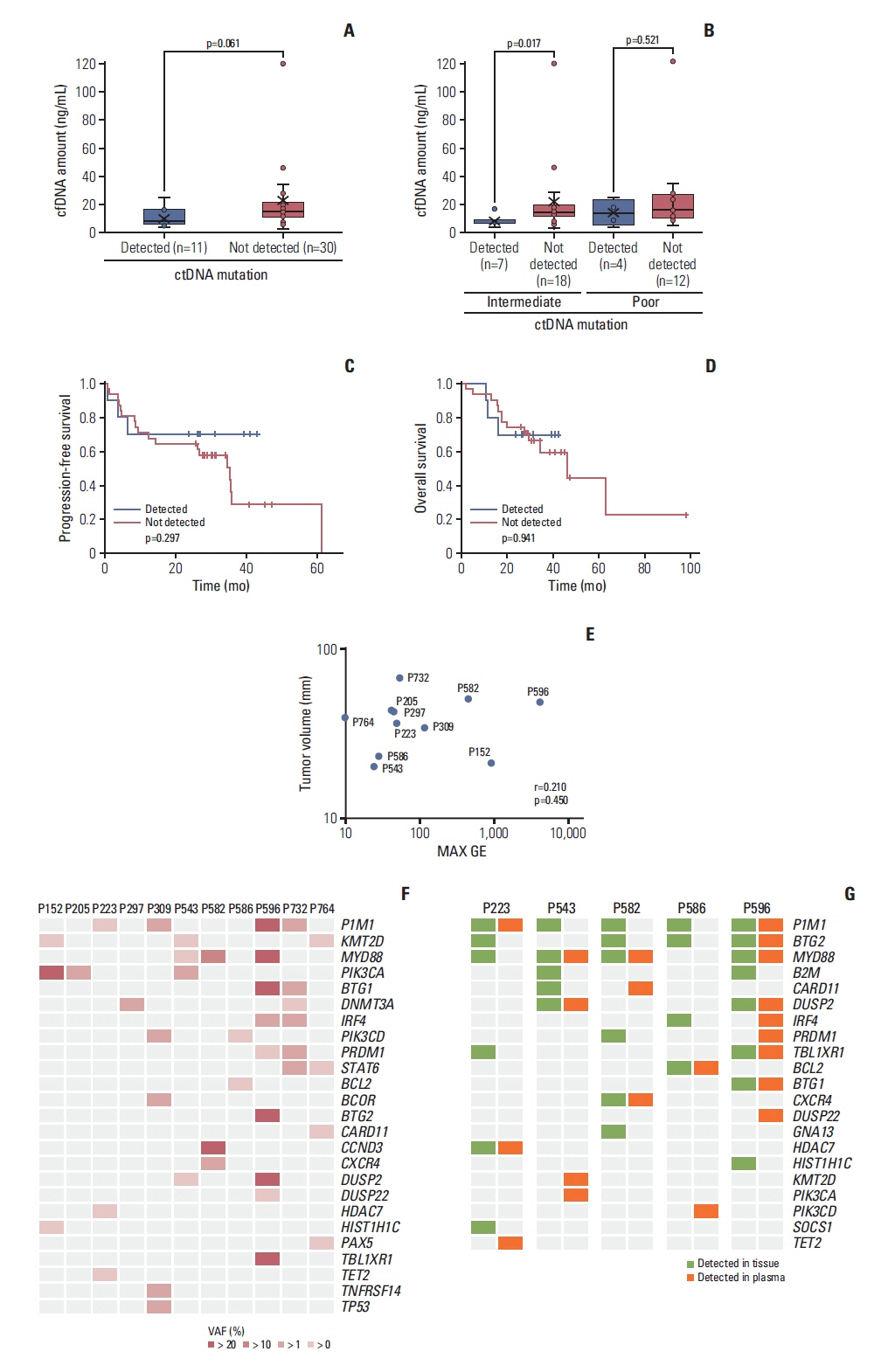
Targeted sequencing of plasma cell-free DNA detected somatic mutations representing ctDNA in 11 cases (11/41, 27%). The detection of ctDNA was not related to the concentration of cell-free DNA or tumor volume. The mutation profiles of these 11 cases varied between patients. The most frequently mutated gene was PIM1 (4/11, 36.4%), whereas KMT2D, PIK3CA, and MYD88 were each observed in three patients (3/11, 27%). The mutations of 13 genes were concordantly found in primary tumor tissue and plasma ctDNA, giving a detection sensitivity of 45%. During the serial tracking of seven patients with complete response, the disappearance of ctDNA mutations was found in four patients, whereas three patients had detected ctDNA mutation at the end of treatment.
Conclusion
The plasma ctDNA mutation analysis still has limited value for surveillance and predicting treatment outcomes of PCNSL because the detection efficiency was lower than other systemic lymphomas. Thus, analytical platforms should be improved to overcome anatomical hurdles associated with PCNSL.
Figure
Cited by 1 articles
-
Feasibility of Circulating Tumor DNA Analysis in Patients with Follicular Lymphoma
Sang Eun Yoon, Seung-Ho Shin, Dae Keun Nam, Junhun Cho, Won Seog Kim, Seok Jin Kim
Cancer Res Treat. 2024;56(3):920-935. doi: 10.4143/crt.2023.869.
Reference
-
References
1. Grommes C, DeAngelis LM. Primary CNS lymphoma. J Clin Oncol. 2017; 35:2410–8.
Article2. Thiel E, Korfel A, Martus P, Kanz L, Griesinger F, Rauch M, et al. High-dose methotrexate with or without whole brain radiotherapy for primary CNS lymphoma (G-PCNSL-SG-1): a phase 3, randomised, non-inferiority trial. Lancet Oncol. 2010; 11:1036–47.
Article3. Hoang-Xuan K, Bessell E, Bromberg J, Hottinger AF, Preusser M, Ruda R, et al. Diagnosis and treatment of primary CNS lymphoma in immunocompetent patients: guidelines from the European Association for Neuro-Oncology. Lancet Oncol. 2015; 16:e322–32.
Article4. Vater I, Montesinos-Rongen M, Schlesner M, Haake A, Purschke F, Sprute R, et al. The mutational pattern of primary lymphoma of the central nervous system determined by whole-exome sequencing. Leukemia. 2015; 29:677–85.
Article5. Zhou Y, Liu W, Xu Z, Zhu H, Xiao D, Su W, et al. Analysis of genomic alteration in primary central nervous system lymphoma and the expression of some related genes. Neoplasia. 2018; 20:1059–69.
Article6. Jen J, Wu L, Sidransky D. An overview on the isolation and analysis of circulating tumor DNA in plasma and serum. Ann N Y Acad Sci. 2000; 906:8–12.
Article7. Gerlinger M, Rowan AJ, Horswell S, Math M, Larkin J, Endesfelder D, et al. Intratumor heterogeneity and branched evolution revealed by multiregion sequencing. N Engl J Med. 2012; 366:883–92.
Article8. Weaver KD, Grossman SA, Herman JG. Methylated tumor-specific DNA as a plasma biomarker in patients with glioma. Cancer Invest. 2006; 24:35–40.
Article9. Liu BL, Cheng JX, Zhang W, Zhang X, Wang R, Lin H, et al. Quantitative detection of multiple gene promoter hypermethylation in tumor tissue, serum, and cerebrospinal fluid predicts prognosis of malignant gliomas. Neuro Oncol. 2010; 12:540–8.
Article10. Best MG, Sol N, Zijl S, Reijneveld JC, Wesseling P, Wurdinger T. Liquid biopsies in patients with diffuse glioma. Acta Neuropathol. 2015; 129:849–65.
Article11. Bettegowda C, Sausen M, Leary RJ, Kinde I, Wang Y, Agrawal N, et al. Detection of circulating tumor DNA in early- and late-stage human malignancies. Sci Transl Med. 2014; 6:224ra24.12. Hickmann AK, Frick M, Hadaschik D, Battke F, Bittl M, Ganslandt O, et al. Molecular tumor analysis and liquid biopsy: a feasibility investigation analyzing circulating tumor DNA in patients with central nervous system lymphomas. BMC Cancer. 2019; 19:192.
Article13. Rossi D, Spina V, Bruscaggin A, Gaidano G. Liquid biopsy in lymphoma. Haematologica. 2019; 104:648–52.
Article14. Hattori K, Sakata-Yanagimoto M, Suehara Y, Yokoyama Y, Kato T, Kurita N, et al. Clinical significance of disease-specific MYD88 mutations in circulating DNA in primary central nervous system lymphoma. Cancer Sci. 2018; 109:225–30.
Article15. Bobillo S, Crespo M, Escudero L, Mayor R, Raheja P, Carpio C, et al. Cell free circulating tumor DNA in cerebrospinal fluid detects and monitors central nervous system involvement of B-cell lymphomas. Haematologica. 2021; 106:513–21.
Article16. Wang X, Gao Y, Shan C, Lai M, He H, Bai B, et al. Association of circulating tumor DNA from the cerebrospinal fluid with high-risk CNS involvement in patients with diffuse large B-cell lymphoma. Clin Transl Med. 2021; 11:e236.
Article17. Abrey LE, Batchelor TT, Ferreri AJ, Gospodarowicz M, Pulczynski EJ, Zucca E, et al. Report of an international workshop to standardize baseline evaluation and response criteria for primary CNS lymphoma. J Clin Oncol. 2005; 23:5034–43.
Article18. Newman AM, Lovejoy AF, Klass DM, Kurtz DM, Chabon JJ, Scherer F, et al. Integrated digital error suppression for improved detection of circulating tumor DNA. Nat Biotechnol. 2016; 34:547–55.
Article19. McKenna A, Hanna M, Banks E, Sivachenko A, Cibulskis K, Kernytsky A, et al. The Genome Analysis Toolkit: a MapReduce framework for analyzing next-generation DNA sequencing data. Genome Res. 2010; 20:1297–303.
Article20. Li H. A statistical framework for SNP calling, mutation discovery, association mapping and population genetical parameter estimation from sequencing data. Bioinformatics. 2011; 27:2987–93.
Article21. Shin SH, Kim YJ, Lee D, Cho D, Ko YH, Cho J, et al. Analysis of circulating tumor DNA by targeted ultra-deep sequencing across various non-Hodgkin lymphoma subtypes. Leuk Lymphoma. 2019; 60:2237–46.
Article22. Fontanilles M, Marguet F, Bohers E, Viailly PJ, Bertrand P, Dubois S, et al. Somatic mutations detected in plasma cell-free DNA by targeted sequencing: assessment of liquid biopsy in primary central nervous system lymphoma. Blood. 2015; 126:332.
Article23. Watanabe J, Natsumeda M, Kanemaru Y, Okada M, Oishi M, Kakita A, et al. Comparison of circulating tumor DNA between body fluids in patients with primary central nervous system lymphoma. Leuk Lymphoma. 2019; 60:3587–9.
Article24. Asmar F, Punj V, Christensen J, Pedersen MT, Pedersen A, Nielsen AB, et al. Genome-wide profiling identifies a DNA methylation signature that associates with TET2 mutations in diffuse large B-cell lymphoma. Haematologica. 2013; 98:1912–20.
Article25. Reddy A, Zhang J, Davis NS, Moffitt AB, Love CL, Waldrop A, et al. Genetic and functional drivers of diffuse large B cell lymphoma. Cell. 2017; 171:481–94.
Article26. Kubuki Y, Yamaji T, Hidaka T, Kameda T, Shide K, Sekine M, et al. TET2 mutation in diffuse large B-cell lymphoma. J Clin Exp Hematop. 2017; 56:145–9.27. Dominguez PM, Ghamlouch H, Rosikiewicz W, Kumar P, Beguelin W, Fontan L, et al. TET2 deficiency causes germinal center hyperplasia, impairs plasma cell differentiation, and promotes B-cell lmphomagenesis. Cancer Discov. 2018; 8:1632–53.28. Fontanilles M, Marguet F, Bohers E, Viailly PJ, Dubois S, Bertrand P, et al. Non-invasive detection of somatic mutations using next-generation sequencing in primary central nervous system lymphoma. Oncotarget. 2017; 8:48157–68.
Article
- Full Text Links
- Actions
-
Cited
- CITED
-
- Close
- Share
- Similar articles
-
- Primary Central Nervous System Lymphoma Mimicking Behcet's Disease
- Primary Non-Hodgkin ''s Lymphoma Involving the Third Ventricle: A Case Report
- Clinical Application of Circulating Tumor DNA Analysis
- Experience of Childhood Non-Hodgkin Lymphoma with Central Nervous System Involvement at Diagnosis
- Primary Central Nervous System Involvement in Peripheral T-Cell Lymphoma: A Case Report